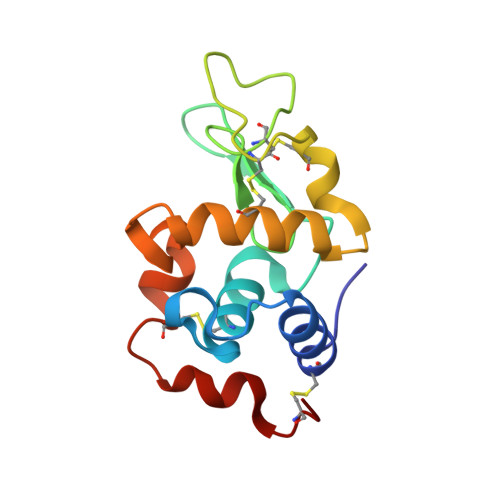
Protein Recognition of Gold-Based Drugs: 3D Structure of the Complex Formed When Lysozyme Reacts with Aubipy(c.).
Messori, L., Cinellu, M.A., Merlino, A.(2014) ACS Med Chem Lett 5: 1110-1113
- PubMed: 25313321 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml500231b
- Primary Citation Related Structures:
4OOT - PubMed Abstract:
The structure of the adduct formed in the reaction between Aubipy(c), a cytotoxic organogold(III) compound, and the model protein hen egg white lysozyme (HEWL) has been solved by X-ray crystallography. It emerges that Aubipy(c), after interaction with HEWL, undergoes reduction of the gold(III) center followed by detaching of the cyclometalated ligand; the resulting naked gold(I) ion is found bound to the protein at Gln121. A direct comparison between the present structure and those previously solved for the lysozyme adducts with other gold(III) compounds demonstrates that coordinated ligands play a key role in the protein-metallodrug recognition process. Structural data support the view that gold(III)-based antitumor prodrugs are activated through metal reduction.
- Department of Chemistry, University of Florence , Via della Lastruccia, 3-13, Florence I-50059, Italy.
Organizational Affiliation: