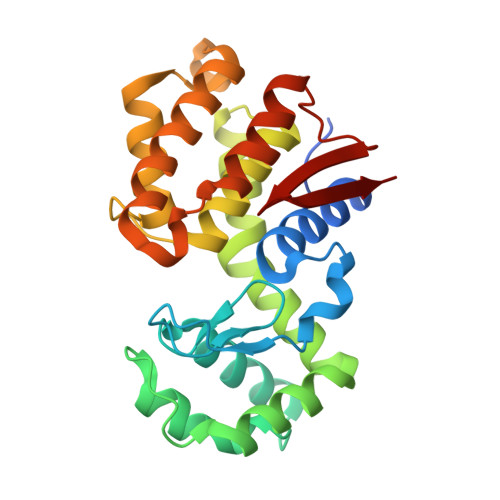
Structural insights into the substrate-binding mechanism for a novel chitosanase.
Lyu, Q., Wang, S., Xu, W., Han, B., Liu, W., Jones, D.N., Liu, W.(2014) Biochem J 461: 335-345
- PubMed: 24766439 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20140159
- Primary Citation Related Structures:
4OLT - PubMed Abstract:
Chitosanase is able to specifically cleave β-1,4-glycosidic bond linkages in chitosan to produce a chito-oligomer product, which has found a variety of applications in many areas, including functional food and cancer therapy. Although several structures for chitosanase have been determined, the substrate-binding mechanism for this enzyme has not been fully elucidated because of the lack of a high-resolution structure of the chitosanase-substrate complex. In the present study we show the crystal structure of a novel chitosanase OU01 from Microbacterium sp. in complex with its substrate hexa-glucosamine (GlcN)6, which belongs to the GH46 (glycoside hydrolyase 46) family in the Carbohydrate Active Enzymes database (http://www.cazy.org/). This structure allows precise determination of the substrate-binding mechanism for the first time. The chitosanase-(GlcN)6 complex structure demonstrates that, from the -2 to +1 position of the (GlcN)6 substrate, the pyranose rings form extensive interactions with the chitosanase-binding cleft. Several residues (Ser27, Tyr37, Arg45, Thr58, Asp60, His203 and Asp235) in the binding cleft are found to form important interactions required to bind the substrate. Site-directed mutagenesis of these residues showed that mutations of Y37F and H203A abolish catalytic activity. In contrast, the mutations T58A and D235A only lead to a moderate loss of catalytic activity, whereas the S27A mutation retains ~80% of the enzymatic activity. In combination with previous mutagenesis studies, these results suggest that the -2, -1 and +1 subsites play a dominant role in substrate binding and catalysis. DSF (differential scanning fluorimetry) assays confirmed that these mutations had no significant effect on protein stability. Taken together, we present the first mechanistic interpretation for the substrate (GlcN)6 binding to chitosanase, which is critical for the design of novel chitosanase used for biomass conversion.
- *College of Marine Life Sciences, Ocean University of China, Qingdao, China.
Organizational Affiliation: