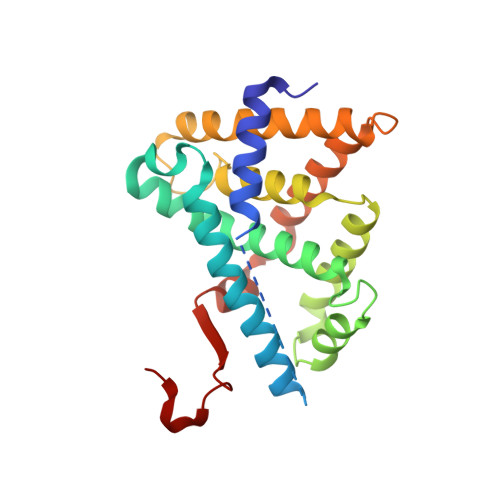
Structural Basis for Small Molecule NDB (N-Benzyl-N-(3-(tert-butyl)-4-hydroxyphenyl)-2,6-dichloro-4-(dimethylamino) Benzamide) as a Selective Antagonist of Farnesoid X Receptor alpha (FXR alpha ) in Stabilizing the Homodimerization of the Receptor.
Xu, X., Xu, X., Liu, P., Zhu, Z.Y., Chen, J., Fu, H.A., Chen, L.L., Hu, L.H., Shen, X.(2015) J Biological Chem 290: 19888-19899
- PubMed: 26100621 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.630475
- Primary Citation Related Structures:
4OIV - PubMed Abstract:
Farnesoid X receptor α (FXRα) as a bile acid sensor plays potent roles in multiple metabolic processes, and its antagonist has recently revealed special interests in the treatment of metabolic disorders, although the underlying mechanisms still remain unclear. Here, we identified that the small molecule N-benzyl-N-(3-(tert-butyl)-4-hydroxyphenyl)-2,6-dichloro-4-(dimethylamino) benzamide (NDB) functioned as a selective antagonist of human FXRα (hFXRα), and the crystal structure of hFXRα ligand binding domain (hFXRα-LBD) in complex with NDB was analyzed. It was unexpectedly discovered that NDB induced rearrangements of helix 11 (H11) and helix 12 (H12, AF-2) by forming a homodimer of hFXRα-LBD, totally different from the active conformation in monomer state, and the binding details were further supported by the mutation analysis. Moreover, functional studies demonstrated that NDB effectively antagonized the GW4064-stimulated FXR/RXR interaction and FXRα target gene expression in primary mouse hepatocytes, including the small heterodimer partner (SHP) and bile-salt export pump (BSEP); meanwhile, administration of NDB to db/db mice efficiently decreased the gene expressions of phosphoenolpyruvate carboxykinase (PEPCK), glucose 6-phosphatase (G6-pase), small heterodimer partner, and BSEP. It is expected that our first analyzed crystal structure of hFXRα-LBD·NDB will help expound the antagonistic mechanism of the receptor, and NDB may find its potential as a lead compound in anti-diabetes research.
- From the CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, 555 Zuchongzhi Road, Shanghai 201203, China, Shanghai Key Laboratory for Bone and Joint Diseases, Shanghai Institute of Traumatology and Orthopaedics, Shanghai Ruijin Hospital, Shanghai Jiaotong University School of Medicine, Shanghai 200025, China.
Organizational Affiliation: