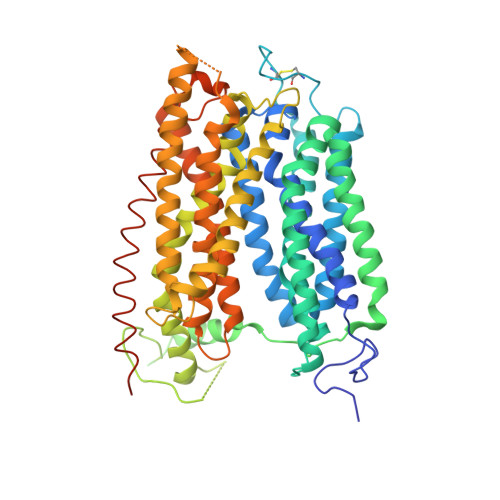
Crystal structure of the plant dual-affinity nitrate transporter NRT1.1.
Sun, J., Bankston, J.R., Payandeh, J., Hinds, T.R., Zagotta, W.N., Zheng, N.(2014) Nature 507: 73-77
- PubMed: 24572362 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13074
- Primary Citation Related Structures:
4OH3 - PubMed Abstract:
Nitrate is a primary nutrient for plant growth, but its levels in soil can fluctuate by several orders of magnitude. Previous studies have identified Arabidopsis NRT1.1 as a dual-affinity nitrate transporter that can take up nitrate over a wide range of concentrations. The mode of action of NRT1.1 is controlled by phosphorylation of a key residue, Thr 101; however, how this post-translational modification switches the transporter between two affinity states remains unclear. Here we report the crystal structure of unphosphorylated NRT1.1, which reveals an unexpected homodimer in the inward-facing conformation. In this low-affinity state, the Thr 101 phosphorylation site is embedded in a pocket immediately adjacent to the dimer interface, linking the phosphorylation status of the transporter to its oligomeric state. Using a cell-based fluorescence resonance energy transfer assay, we show that functional NRT1.1 dimerizes in the cell membrane and that the phosphomimetic mutation of Thr 101 converts the protein into a monophasic high-affinity transporter by structurally decoupling the dimer. Together with analyses of the substrate transport tunnel, our results establish a phosphorylation-controlled dimerization switch that allows NRT1.1 to uptake nitrate with two distinct affinity modes.
- Department of Pharmacology, Box 357280, University of Washington, Seattle, Washington 98195, USA.
Organizational Affiliation: