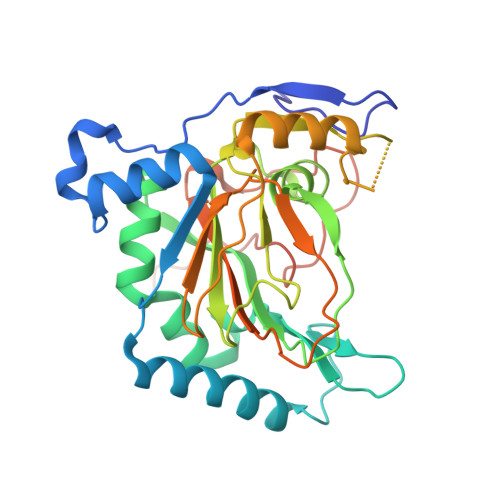
Biochemical properties of ectoine hydroxylases from extremophiles and their wider taxonomic distribution among microorganisms.
Widderich, N., Hoppner, A., Pittelkow, M., Heider, J., Smits, S.H., Bremer, E.(2014) PLoS One 9: e93809-e93809
- PubMed: 24714029 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0093809
- Primary Citation Related Structures:
4NMI - PubMed Abstract:
Ectoine and hydroxyectoine are well-recognized members of the compatible solutes and are widely employed by microorganisms as osmostress protectants. The EctABC enzymes catalyze the synthesis of ectoine from the precursor L-aspartate-β-semialdehyde. A subgroup of the ectoine producers can convert ectoine into 5-hydroxyectoine through a region-selective and stereospecific hydroxylation reaction. This compatible solute possesses stress-protective and function-preserving properties different from those of ectoine. Hydroxylation of ectoine is carried out by the EctD protein, a member of the non-heme-containing iron (II) and 2-oxoglutarate-dependent dioxygenase superfamily. We used the signature enzymes for ectoine (EctC) and hydroxyectoine (EctD) synthesis in database searches to assess the taxonomic distribution of potential ectoine and hydroxyectoine producers. Among 6428 microbial genomes inspected, 440 species are predicted to produce ectoine and of these, 272 are predicted to synthesize hydroxyectoine as well. Ectoine and hydroxyectoine genes are found almost exclusively in Bacteria. The genome context of the ect genes was explored to identify proteins that are functionally associated with the synthesis of ectoines; the specialized aspartokinase Ask_Ect and the regulatory protein EctR. This comprehensive in silico analysis was coupled with the biochemical characterization of ectoine hydroxylases from microorganisms that can colonize habitats with extremes in salinity (Halomonas elongata), pH (Alkalilimnicola ehrlichii, Acidiphilium cryptum), or temperature (Sphingopyxis alaskensis, Paenibacillus lautus) or that produce hydroxyectoine very efficiently over ectoine (Pseudomonas stutzeri). These six ectoine hydroxylases all possess similar kinetic parameters for their substrates but exhibit different temperature stabilities and differ in their tolerance to salts. We also report the crystal structure of the Virgibacillus salexigens EctD protein in its apo-form, thereby revealing that the iron-free structure exists already in a pre-set configuration to incorporate the iron catalyst. Collectively, our work defines the taxonomic distribution and salient biochemical properties of the ectoine hydroxylase protein family and contributes to the understanding of its structure.
- Laboratory for Microbiology, Department of Biology, Philipps-University Marburg, Marburg, Germany; Max Planck Institute for Terrestrial Microbiology, Emeritus Group R.K. Thauer, Marburg, Germany.
Organizational Affiliation: