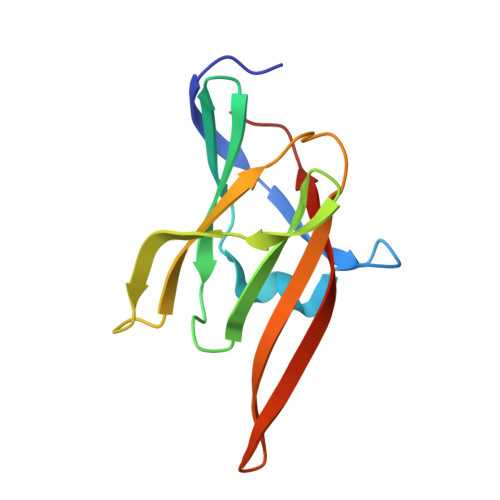
Novel Mechanism of Hemin Capture by Hbp2, the Hemoglobin-binding Hemophore from Listeria monocytogenes.
Malmirchegini, G.R., Sjodt, M., Shnitkind, S., Sawaya, M.R., Rosinski, J., Newton, S.M., Klebba, P.E., Clubb, R.T.(2014) J Biological Chem 289: 34886-34899
- PubMed: 25315777
- DOI: https://doi.org/10.1074/jbc.M114.583013
- Primary Citation Related Structures:
4MYP, 4NLA - PubMed Abstract:
Iron is an essential nutrient that is required for the growth of the bacterial pathogen Listeria monocytogenes. In cell cultures, this microbe secretes hemin/hemoglobin-binding protein 2 (Hbp2; Lmo2185) protein, which has been proposed to function as a hemophore that scavenges heme from the environment. Based on its primary sequence, Hbp2 contains three NEAr transporter (NEAT) domains of unknown function. Here we show that each of these domains mediates high affinity binding to ferric heme (hemin) and that its N- and C-terminal domains interact with hemoglobin (Hb). The results of hemin transfer experiments are consistent with Hbp2 functioning as an Hb-binding hemophore that delivers hemin to other Hbp2 proteins that are attached to the cell wall. Surprisingly, our work reveals that the central NEAT domain in Hbp2 binds hemin even though its primary sequence lacks a highly conserved YXXXY motif that is used by all other previously characterized NEAT domains to coordinate iron in the hemin molecule. To elucidate the mechanism of hemin binding by Hbp2, we determined crystal structures of its central NEAT domain (Hbp2(N2); residues 183-303) in its free and hemin-bound states. The structures reveal an unprecedented mechanism of hemin binding in which Hbp2(N2) undergoes a major conformational rearrangement that facilitates metal coordination by a non-canonical tyrosine residue. These studies highlight previously unrecognized plasticity in the hemin binding mechanism of NEAT domains and provide insight into how L. monocytogenes captures heme iron.
- From the Department of Chemistry and Biochemistry and the UCLA-Department of Energy Institute for Genomics and Proteomics, UCLA, Los Angeles, California 90095 and.
Organizational Affiliation: