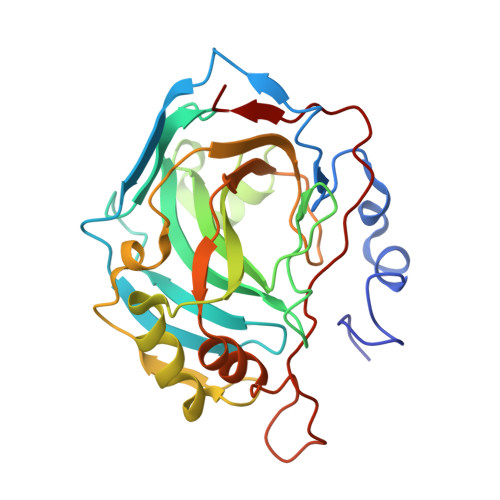
Structure of a complex formed by a protein and a helical aromatic oligoamide foldamer at 2.1 a resolution.
Buratto, J., Colombo, C., Stupfel, M., Dawson, S.J., Dolain, C., Langlois d'Estaintot, B., Fischer, L., Granier, T., Laguerre, M., Gallois, B., Huc, I.(2014) Angew Chem Int Ed Engl 53: 883-887
- PubMed: 24288253 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201309160
- Primary Citation Related Structures:
4LP6, 4MTY - PubMed Abstract:
In the search of molecules that could recognize sizeable areas of protein surfaces, a series of ten helical aromatic oligoamide foldamers was synthesized on solid phase. The foldamers comprise three to five monomers carrying various proteinogenic side chains, and exist as racemic mixtures of interconverting right-handed and left-handed helices. Functionalization of the foldamers by a nanomolar ligand of human carbonic anhydrase II (HCA) ensured that they would be held in close proximity to the protein surface. Foldamer-protein interactions were screened by circular dichroism (CD). One foldamer displayed intense CD bands indicating that a preferred helix handedness is induced upon interacting with the protein surface. The crystal structure of the complex between this foldamer and HCA could be resolved at 2.1 Å resolution and revealed a number of unanticipated protein-foldamer, foldamer-foldamer, and protein-protein interactions.
- Université de Bordeaux, CBMN, UMR 5248, Institut Européen de Chimie Biologie, 2 rue Robert Escarpit 33607 Pessac (France); CNRS, CBMN, UMR5248 (France); Institut Polytechnique de Bordeaux, UMR5248 (France).
Organizational Affiliation: