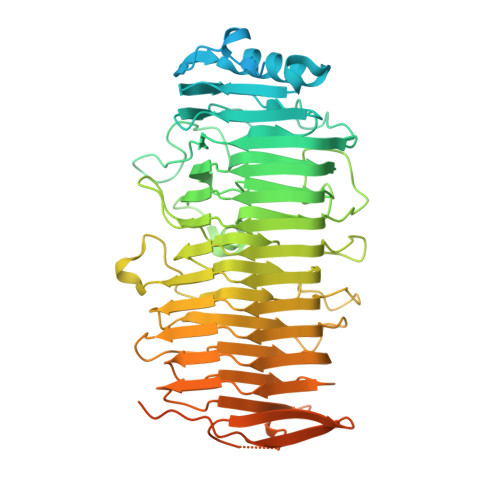
Crystal structure of PfbA, a surface adhesin of Streptococcus pneumoniae, provides hints into its interaction with fibronectin
Beulin, D.S.J., Yamaguchi, M., Kawabata, S., Ponnuraj, K.(2013) Int J Biol Macromol 64C: 168-173
- PubMed: 24321492
- DOI: https://doi.org/10.1016/j.ijbiomac.2013.11.035
- Primary Citation of Related Structures:
4MR0 - PubMed Abstract:
PfbA is a surface adhesin and invasin of Streptococcus pneumoniae that binds to human fibronectin and plasminogen of the host extracellular matrix. It is a virulence factor for its pathogenesis. The crystal structure of recombinant PfbA150-607 from S. pneumoniae strain R6, was determined using multiwavelength anomalous dispersion (MAD) method and refined to 1.90Å resolution. The structure of rPfbA150-607 revealed that residues Thr150 to Lys570 form a rigid parallel beta helix, followed by a short disordered region (571-607) that consists of beta hairpins. The structural organization of the beta helix resembles that of polysaccharide-modifying enzymes. The structural and sequence features essential for fibronectin-binding observed in the well characterized fibronectin-binding proteins such as FnBPA of Staphylococcus aureus, SfbI of Streptococcus pyogenes and BBK32 of Borrelia burgdorferi has been found in rPfbA150-607. Based on this, it is predicted that the disordered region following the beta helix could be the fibronectin-binding region in PfbA. PfbA150-607 contains relatively high number of surface exposed lysines and these residues are probably involved in binding plasmin(ogen) as observed in other plasminogen-binding proteins.
- Centre of Advanced Study in Crystallography and Biophysics, University of Madras, Guindy Campus, Chennai 600 025, India.
Organizational Affiliation: