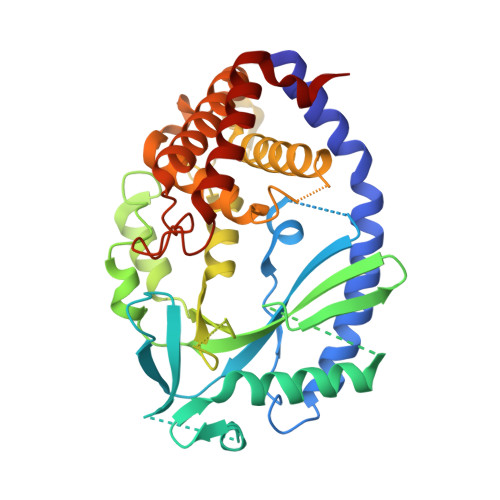
Structural and Functional Analyses of DNA-Sensing and Immune Activation by Human cGAS
Kato, K., Ishii, R., Goto, E., Ishitani, R., Tokunaga, F., Nureki, O.(2013) PLoS One 8: e76983-e76983
- PubMed: 24116191
- DOI: https://doi.org/10.1371/journal.pone.0076983
- Primary Citation Related Structures:
4MKP - PubMed Abstract:
The detection of cytosolic DNA, derived from pathogens or host cells, by cytosolic receptors is essential for appropriate host immune responses. Cyclic GMP-AMP synthase (cGAS) is a newly identified cytosolic DNA receptor that produces cyclic GMP-AMP, which activates stimulator of interferon genes (STING), resulting in TBK1-IRF3 pathway activation followed by the production of type I interferons. Here we report the crystal structure of human cGAS. The structure revealed that a cluster of lysine and arginine residues forms the positively charged DNA binding surface of human cGAS, which is important for the STING-dependent immune activation. A structural comparison with other previously determined cGASs and our functional analyses suggested that a conserved zinc finger motif and a leucine residue on the DNA binding surface are crucial for the DNA-specific immune response of human cGAS, consistent with previous work. These structural features properly orient the DNA binding to cGAS, which is critical for DNA-induced cGAS activation and STING-dependent immune activation. Furthermore, we showed that the cGAS-induced activation of STING also involves the activation of the NF-κB and IRF3 pathways. Our results indicated that cGAS is a DNA sensor that efficiently activates the host immune system by inducing two distinct pathways.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Bunkyo-ku, Tokyo, Japan ; Global Research Cluster, RIKEN, Wako, Saitama, Japan.
Organizational Affiliation: