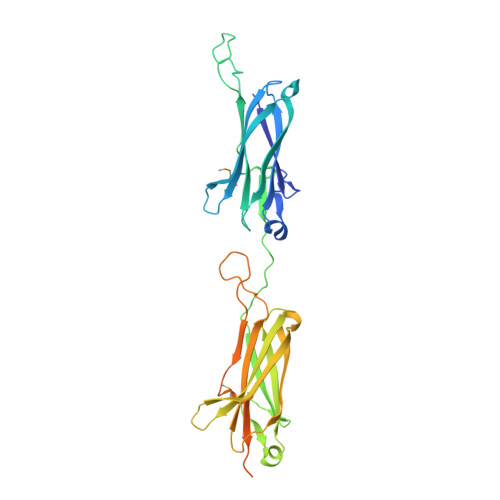
Autocatalytically generated Thr-Gln ester bond cross-links stabilize the repetitive Ig-domain shaft of a bacterial cell surface adhesin.
Kwon, H., Squire, C.J., Young, P.G., Baker, E.N.(2014) Proc Natl Acad Sci U S A 111: 1367-1372
- PubMed: 24344302
- DOI: https://doi.org/10.1073/pnas.1316855111
- Primary Citation Related Structures:
4MKM, 4NI6 - PubMed Abstract:
Gram-positive bacteria are decorated by a variety of proteins that are anchored to the cell wall and project from it to mediate colonization, attachment to host cells, and pathogenesis. These proteins, and protein assemblies, such as pili, are typically long and thin yet must withstand high levels of mechanical stress and proteolytic attack. The recent discovery of intramolecular isopeptide bond cross-links, formed autocatalytically, in the pili from Streptococcus pyogenes has highlighted the role that such cross-links can play in stabilizing such structures. We have investigated a putative cell-surface adhesin from Clostridium perfringens comprising an N-terminal adhesin domain followed by 11 repeat domains. The crystal structure of a two-domain fragment shows that each domain has an IgG-like fold and contains an unprecedented ester bond joining Thr and Gln side chains. MS confirms the presence of these bonds. We show that the bonds form through an autocatalytic intramolecular reaction catalyzed by an adjacent His residue in a serine protease-like mechanism. Two buried acidic residues assist in the reaction. By mutagenesis, we show that loss of the ester bond reduces the thermal stability drastically and increases susceptibility to proteolysis. As in pilin domains, the bonds are placed at a strategic position joining the first and last strands, even though the Ig fold type differs. Bioinformatic analysis suggests that similar domains and ester bond cross-links are widespread in Gram-positive bacterial adhesins.
- Maurice Wilkins Centre for Molecular Biodiscovery and School of Biological Sciences, University of Auckland, Auckland 1010, New Zealand.
Organizational Affiliation: