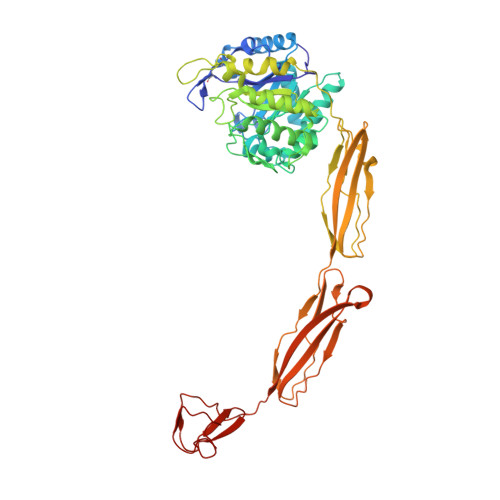
Crystal structures of substrate-bound chitinase from the psychrophilic bacterium Moritella marina and its structure in solution
Malecki, P.H., Vorgias, C.E., Petoukhov, M.V., Svergun, D.I., Rypniewski, W.(2014) Acta Crystallogr D Biol Crystallogr 70: 676-684
- PubMed: 24598737
- DOI: https://doi.org/10.1107/S1399004713032264
- Primary Citation Related Structures:
4MB3, 4MB4, 4MB5 - PubMed Abstract:
The four-domain structure of chitinase 60 from Moritella marina (MmChi60) is outstanding in its complexity. Many glycoside hydrolases, such as chitinases and cellulases, have multi-domain structures, but only a few have been solved. The flexibility of the hinge regions between the domains apparently makes these proteins difficult to crystallize. The analysis of an active-site mutant of MmChi60 in an unliganded form and in complex with the substrates NAG4 and NAG5 revealed significant differences in the substrate-binding site compared with the previously determined complexes of most studied chitinases. A SAXS experiment demonstrated that in addition to the elongated state found in the crystal, the protein can adapt other conformations in solution ranging from fully extended to compact.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Noskowskiego 12/14, 61-704 Poznan, Poland.
Organizational Affiliation: