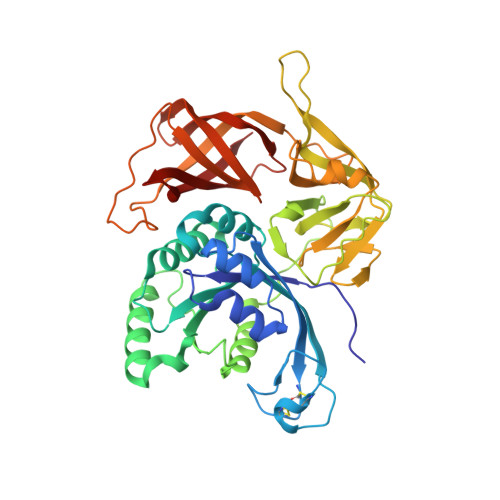
Conformational transitions in the gamma subunit of the archaeal translation initiation factor 2.
Nikonov, O., Stolboushkina, E., Arkhipova, V., Kravchenko, O., Nikonov, S., Garber, M.(2014) Acta Crystallogr D Biol Crystallogr 70: 658-667
- PubMed: 24598735
- DOI: https://doi.org/10.1107/S1399004713032240
- Primary Citation Related Structures:
4M0L, 4M2L, 4M4S, 4M53 - PubMed Abstract:
In eukaryotes and archaea, the heterotrimeric translation initiation factor 2 (e/aIF2) is pivotal for the delivery of methionylated initiator tRNA (Met-tRNA(i)) to the ribosome. It acts as a molecular switch that cycles between inactive (GDP-bound) and active (GTP-bound) states. Recent studies show that eIF2 can also exist in a long-lived eIF2γ-GDP-P(i) (inorganic phosphate) active state. Here, four high-resolution crystal structures of aIF2γ from Sulfolobus solfataricus are reported: aIF2γ-GDPCP (a nonhydrolyzable GTP analogue), aIF2γ-GDP-formate (in which a formate ion possibly mimics P(i)), aIF2γ-GDP and nucleotide-free aIF2γ. The structures describe the different states of aIF2γ and demonstrate the conformational transitions that take place in the aIF2γ `life cycle'.
- Institute of Protein Research, Russian Academy of Sciences, Pushchino 142290, Moscow Region, Russian Federation.
Organizational Affiliation: