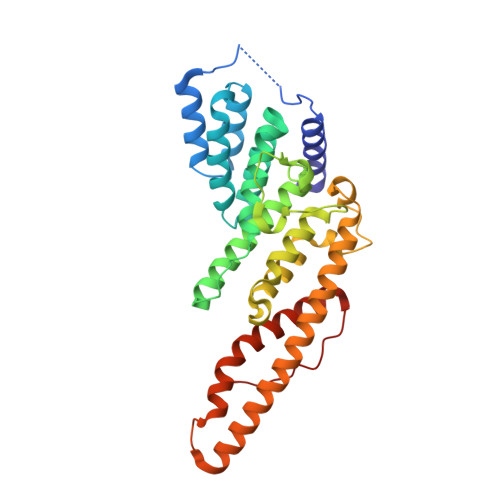
A highly conserved region essential for NMD in the Upf2 N-terminal domain.
Fourati, Z., Roy, B., Millan, C., Coureux, P.D., Kervestin, S., van Tilbeurgh, H., He, F., Uson, I., Jacobson, A., Graille, M.(2014) J Mol Biology 426: 3689-3702
- PubMed: 25277656 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2014.09.015
- Primary Citation Related Structures:
4LUN - PubMed Abstract:
Upf1, Upf2, and Upf3 are the principal regulators of nonsense-mediated mRNA decay (NMD), a cytoplasmic surveillance pathway that accelerates the degradation of mRNAs undergoing premature translation termination. These three proteins interact with each other, the ribosome, the translation termination machinery, and multiple mRNA decay factors, but the precise mechanism allowing the selective detection and degradation of nonsense-containing transcripts remains elusive. Here, we have determined the crystal structure of the N-terminal mIF4G domain from Saccharomyces cerevisiae Upf2 and identified a highly conserved region in this domain that is essential for NMD and independent of Upf2's binding sites for Upf1 and Upf3. Mutations within this conserved region not only inactivate NMD but also disrupt Upf2 binding to specific proteins, including Dbp6, a DEAD-box helicase. Although current models indicate that Upf2 functions principally as an activator of Upf1 and a bridge between Upf1 and Upf3, our data suggest that it may also serve as a platform for the association of additional factors that play roles in premature translation termination and NMD.
- Laboratoire de Biochimie, CNRS UMR 7654, Ecole Polytechnique, F-91128 Palaiseau Cedex, France; Institut de Biochimie et Biophysique Moléculaire et Cellulaire, CNRS UMR 8619, Université Paris Sud, F-91405 Orsay Cedex, France.
Organizational Affiliation: