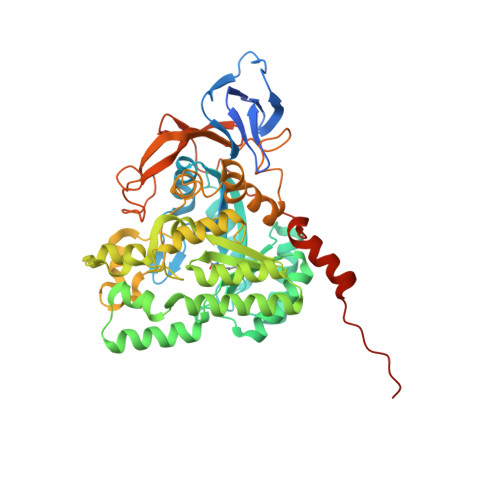
Crystal structures of vertebrate dihydropyrimidinase and complexes from Tetraodon nigroviridis with lysine carbamylation: metal and structural requirements for post-translational modification and function.
Hsieh, Y.C., Chen, M.C., Hsu, C.C., Chan, S.I., Yang, Y.S., Chen, C.J.(2013) J Biological Chem 288: 30645-30658
- PubMed: 24005677
- DOI: https://doi.org/10.1074/jbc.M113.496778
- Primary Citation Related Structures:
4LCQ, 4LCR, 4LCS - PubMed Abstract:
Lysine carbamylation, a post-translational modification, facilitates metal coordination for specific enzymatic activities. We have determined structures of the vertebrate dihydropyrimidinase from Tetraodon nigroviridis (TnDhp) in various states: the apoenzyme as well as two forms of the holoenzyme with one and two metals at the catalytic site. The essential active-site structural requirements have been identified for the possible existence of four metal-mediated stages of lysine carbamylation. Only one metal is sufficient for stabilizing lysine carbamylation; however, the post-translational lysine carbamylation facilitates additional metal coordination for the regulation of specific enzymatic activities through controlling the conformations of two dynamic loops, Ala(69)-Arg(74) and Met(158)-Met(165), located in the tunnel for the substrate entrance. The substrate/product tunnel is in the "open form" in the apo-TnDhp, in the "intermediate state" in the monometal TnDhp, and in the "closed form" in the dimetal TnDhp structure, respectively. Structural comparison also suggests that the C-terminal tail plays a role in the enzymatic function through interactions with the Ala(69)-Arg(74) dynamic loop. In addition, the structures of the dimetal TnDhp in complexes with hydantoin, N-carbamyl-β-alanine, and N-carbamyl-β-amino isobutyrate as well as apo-TnDhp in complex with a product analog, N-(2-acetamido)-iminodiacetic acid, have been determined. These structural results illustrate how a protein exploits unique lysines and the metal distribution to accomplish lysine carbamylation as well as subsequent enzymatic functions.
- From the Life Science Group, Scientific Research Division, National Synchrotron Radiation Research Center, Hsinchu 30076, Taiwan.
Organizational Affiliation: