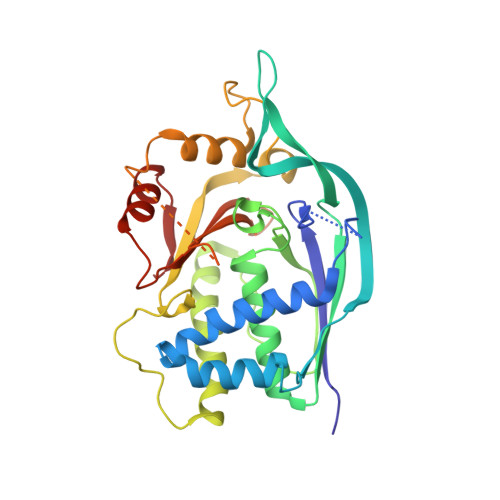
Crystal structure and CRISPR RNA-binding site of the Cmr1 subunit of the Cmr interference complex
Sun, J., Jeon, J.H., Shin, M., Shin, H.C., Oh, B.H., Kim, J.S.(2014) Acta Crystallogr D Biol Crystallogr 70: 535-543
- PubMed: 24531487
- DOI: https://doi.org/10.1107/S1399004713030290
- Primary Citation of Related Structures:
4L6U - PubMed Abstract:
A multi-subunit ribonucleoprotein complex termed the Cmr RNA-silencing complex recognizes and destroys viral RNA in the CRISPR-mediated immune defence mechanism in many prokaryotes using an as yet unclear mechanism. In Archaeoglobus fulgidus, this complex consists of six subunits, Cmr1-Cmr6. Here, the crystal structure of Cmr1 from A. fulgidus is reported, revealing that the protein is composed of two tightly associated ferredoxin-like domains. The domain located at the N-terminus is structurally most similar to the N-terminal ferredoxin-like domain of the CRISPR RNA-processing enzyme Cas6 from Pyrococcus furiosus. An ensuing mutational analysis identified a highly conserved basic surface patch that binds single-stranded nucleic acids specifically, including the mature CRISPR RNA, but in a sequence-independent manner. In addition, this subunit was found to cleave single-stranded RNA. Together, these studies elucidate the structure and the catalytic activity of the Cmr1 subunit.
- Department of Chemistry, Chonnam National University, Gwangju 500-757, Republic of Korea.
Organizational Affiliation: