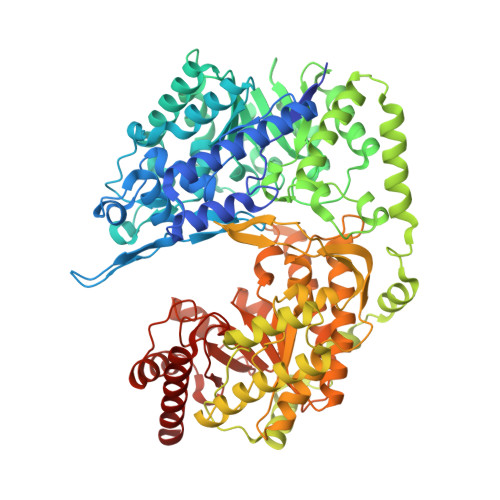
Structural analysis of a fungal methionine synthase with substrates and inhibitors.
Ubhi, D., Kago, G., Monzingo, A.F., Robertus, J.D.(2014) J Mol Biology 426: 1839-1847
- PubMed: 24524835 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2014.02.006
- Primary Citation Related Structures:
4L5Z, 4L61, 4L64, 4L65, 4L6H, 4L6O - PubMed Abstract:
The cobalamin-independent methionine synthase from Candida albicans, known as Met6p, is a 90-kDa enzyme that consists of two (βα)8 barrels. The active site is located between the two domains and has binding sites for a zinc ion and substrates L-homocysteine and 5-methyl-tetrahydrofolate-glutamate3. Met6p catalyzes transfer of the methyl group of 5-methyl-tetrahydrofolate-glutamate3 to the L-homocysteine thiolate to generate methionine. Met6p is essential for fungal growth, and we currently pursue it as an antifungal drug design target. Here we report the binding of L-homocysteine, methionine, and several folate analogs. We show that binding of L-homocysteine or methionine results in conformational rearrangements at the amino acid binding pocket, moving the catalytic zinc into position to activate the thiol group. We also map the folate binding pocket and identify specific binding residues, like Asn126, whose mutation eliminates catalytic activity. We also report the development of a robust fluorescence-based activity assay suitable for high-throughput screening. We use this assay and an X-ray structure to characterize methotrexate as a weak inhibitor of fungal Met6p.
- Institute for Cellular and Molecular Biology, Department of Molecular Biosciences, University of Texas at Austin, Austin, TX 78712, USA.
Organizational Affiliation: