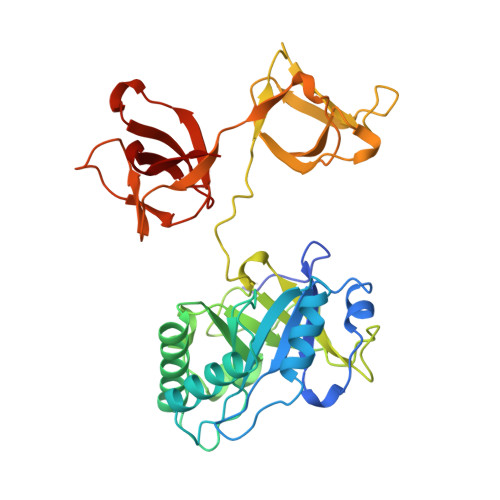
X-ray structure of a novel endolysin encoded by episomal phage phiSM101 of Clostridium perfringens.
Tamai, E., Yoshida, H., Sekiya, H., Nariya, H., Miyata, S., Okabe, A., Kuwahara, T., Maki, J., Kamitori, S.(2014) Mol Microbiol 92: 326-337
- PubMed: 24674022 Search on PubMed
- DOI: https://doi.org/10.1111/mmi.12559
- Primary Citation Related Structures:
4KRT, 4KRU - PubMed Abstract:
Gram-positive bacteria possess a thick cell wall composed of a mesh polymer of peptidoglycans, which provides physical protection. Endolysins encoded by phages infecting bacteria can hydrolyse peptidoglycans in the bacterial cell wall, killing the host bacteria immediately. The endolysin (Psm) encoded by episomal phage phiSM101 of enterotoxigenic Clostridium perfringens type A strain SM101 exhibits potent lytic activity towards most strains of Clostridium perfringens. Psm has an N-terminal catalytic domain highly homologous to N-acetylmuramidases belonging to the glycoside hydrolase 25 family, and C-terminal tandem repeated bacterial Src homology 3 (SH3_3) domains as the cell wall-binding domain. The X-ray structure of full-length Psm and a catalytic domain of Psm in complex with N-acetylglucosamine were determined to elucidate the catalytic reaction and cell wall recognition mechanisms of Psm. The results showed that Psm may have adopted a neighbouring-group mechanism for the catalytic hydrolysing reaction in which the N-acetyl carbonyl group of the substrate was involved in the formation of an oxazolinium ion intermediate. Based on structural comparisons with other endolysins and a modelling study, we proposed that tandem repeated SH3_3 domains of Psm recognized the peptide side-chains of peptidoglycans to assist the catalytic domain hydrolysing the glycan backbone.
- Life Science Research Center, Kagawa University, 1750-1, Ikenobe, Miki-cho, Kita-gun, Kagawa, 761-0793, Japan; Department of Infectious Disease, College of Pharmaceutical Science, Matsuyama University, 4-2 Bunkyo-cho, Matsuyama, Ehime, 790-8578, Japan.
Organizational Affiliation: