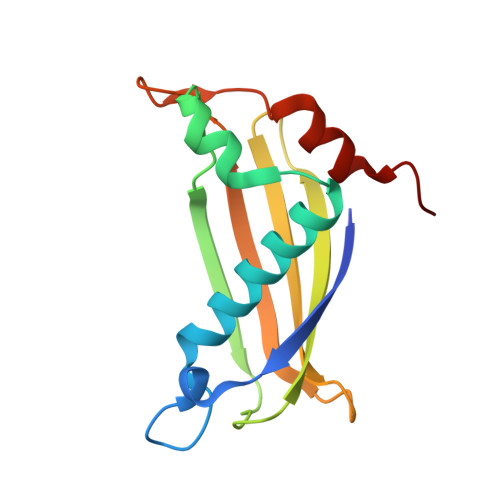
Functional convergence of structurally distinct thioesterases from cyanobacteria and plants involved in phylloquinone biosynthesis.
Furt, F., Allen, W.J., Widhalm, J.R., Madzelan, P., Rizzo, R.C., Basset, G., Wilson, M.A.(2013) Acta Crystallogr D Biol Crystallogr 69: 1876-1888
- PubMed: 24100308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913015771
- Primary Citation Related Structures:
4K00, 4K02 - PubMed Abstract:
The synthesis of phylloquinone (vitamin K1) in photosynthetic organisms requires a thioesterase that hydrolyzes 1,4-dihydroxy-2-naphthoyl-CoA (DHNA-CoA) to release 1,4-dihydroxy-2-naphthoate (DHNA). Cyanobacteria and plants contain distantly related hotdog-fold thioesterases that catalyze this reaction, although the structural basis of these convergent enzymatic activities is unknown. To investigate this, the crystal structures of hotdog-fold DHNA-CoA thioesterases from the cyanobacterium Synechocystis (Slr0204) and the flowering plant Arabidopsis thaliana (AtDHNAT1) were determined. These enzymes form distinct homotetramers and use different active sites to catalyze hydrolysis of DHNA-CoA, similar to the 4-hydroxybenzoyl-CoA (4-HBA-CoA) thioesterases from Pseudomonas and Arthrobacter. Like the 4-HBA-CoA thioesterases, the DHNA-CoA thioesterases contain either an active-site aspartate (Slr0204) or glutamate (AtDHNAT1) that are predicted to be catalytically important. Computational modeling of the substrate-bound forms of both enzymes indicates the residues that are likely to be involved in substrate binding and catalysis. Both enzymes are selective for DHNA-CoA as a substrate, but this selectivity is achieved using divergent predicted binding strategies. The Slr0204 binding pocket is predominantly hydrophobic and closely conforms to DHNA, while that of AtDHNAT1 is more polar and solvent-exposed. Considered in light of the related 4-HBA-CoA thioesterases, these structures indicate that hotdog-fold thioesterases using either an active-site aspartate or glutamate diverged into distinct clades prior to the evolution of strong substrate specificity in these enzymes.
- Center for Plant Science Innovation and Departments of Agronomy and Horticulture and Department of Biochemistry, University of Nebraska, Lincoln, NE 68588, USA.
Organizational Affiliation: