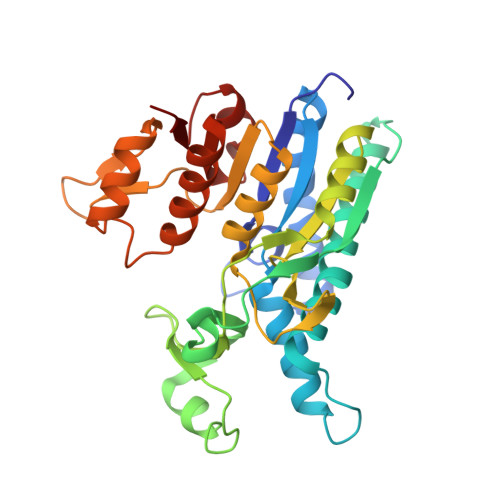
Crystal Structures of Carbamate Kinase from Giardia lamblia Bound with Citric Acid and AMP-PNP.
Lim, K., Kulakova, L., Galkin, A., Herzberg, O.(2013) PLoS One 8: e64004-e64004
- PubMed: 23700444
- DOI: https://doi.org/10.1371/journal.pone.0064004
- Primary Citation of Related Structures:
4JZ7, 4JZ8, 4JZ9 - PubMed Abstract:
The parasite Giardia lamblia utilizes the L-arginine dihydrolase pathway to generate ATP from L-arginine. Carbamate kinase (CK) catalyzes the last step in this pathway, converting ADP and carbamoyl phosphate to ATP and ammonium carbamate. Because the L-arginine pathway is essential for G. lamblia survival and absent in high eukaryotes including humans, the enzyme is a potential target for drug development. We have determined two crystal structures of G. lamblia CK (glCK) with bound ligands. One structure, in complex with a nonhydrolyzable ATP analog, adenosine 5'-adenylyl-β,γ-imidodiphosphate (AMP-PNP), was determined at 2.6 Å resolution. The second structure, in complex with citric acid bound in the postulated carbamoyl phosphate binding site, was determined in two slightly different states at 2.1 and 2.4 Å resolution. These structures reveal conformational flexibility of an auxiliary domain (amino acid residues 123-170), which exhibits open or closed conformations or structural disorder, depending on the bound ligand. The structures also reveal a smaller conformational change in a region associated the AMP-PNP adenine binding site. The protein residues involved in binding, together with a model of the transition state, suggest that catalysis follows an in-line, predominantly dissociative, phosphotransfer reaction mechanism, and that closure of the flexible auxiliary domain is required to protect the transition state from bulk solvent.
- Institute for Bioscience and Biotechnology Research, University of Maryland, Rockville, Maryland, USA.
Organizational Affiliation: