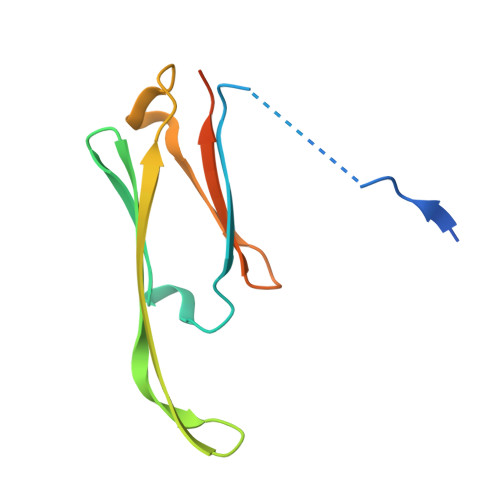
Molecular structure and dynamics of the dimeric human small heat shock protein HSPB6.
Weeks, S.D., Baranova, E.V., Heirbaut, M., Beelen, S., Shkumatov, A.V., Gusev, N.B., Strelkov, S.V.(2014) J Struct Biol 185: 342-354
- PubMed: 24382496 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2013.12.009
- Primary Citation Related Structures:
4JUS, 4JUT - PubMed Abstract:
ATP-independent small heat-shock proteins (sHSPs) are an essential component of the cellular chaperoning machinery. Under both normal and stress conditions, sHSPs bind partially unfolded proteins and prevent their irreversible aggregation. Canonical vertebrate sHSPs, such as the α-crystallins, form large polydisperse oligomers from which smaller, functionally active subspecies dissociate. Here we focus on human HSPB6 which, despite having considerable homology to the α-crystallins in both the N-terminal region and the signature α-crystallin domain (ACD), only forms dimers in solution that represent the basic chaperoning subspecies. We addressed the three-dimensional structure and functional properties of HSPB6 in a hybrid study employing X-ray crystallography, solution small-angle X-ray scattering (SAXS), mutagenesis, size-exclusion chromatography and chaperoning assays. The crystal structure of a proteolytically stable fragment reveals typical ACD dimers which further form tetrameric assemblies as a result of extensive inter-dimer patching of the β4/β8 grooves. The patching is surprisingly mediated by tripeptide motifs, found in the N-terminal domain directly adjacent to the ACD, that are resembling but distinct from the canonical IxI sequence commonly binding this groove. By combining the crystal structure with SAXS data for the full-length protein, we derive a molecular model of the latter. In solution, HSPB6 shows a strong attractive self-interaction, a property that correlates with its chaperoning activity. Both properties are dictated by the unstructured yet compact N-terminal domain, specifically a region highly conserved across vertebrate sHSPs.
- Laboratory for Biocrystallography, Department of Pharmaceutical and Pharmacological Sciences, KU Leuven, Belgium.
Organizational Affiliation: