Structural basis for interaction between Mycobacterium smegmatis Ms6564, a TetR family master regulator, and its target DNA.
Yang, S., Gao, Z., Li, T., Yang, M., Zhang, T., Dong, Y., He, Z.G.(2013) J Biological Chem 288: 23687-23695
- PubMed: 23803605 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.468694
- Primary Citation Related Structures:
4JKZ, 4JL3 - PubMed Abstract:
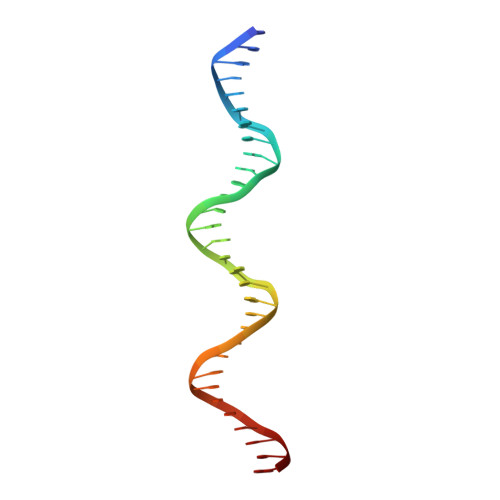
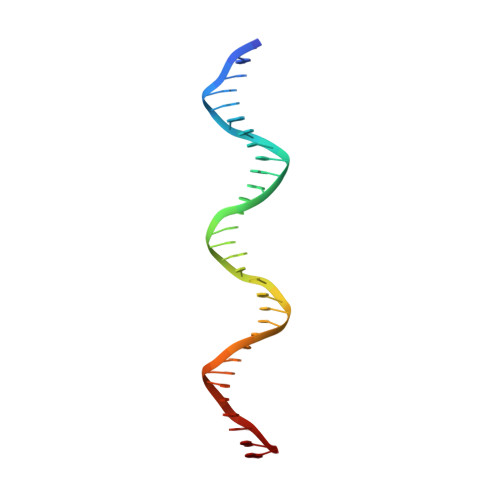
Master regulators, which broadly affect expression of diverse genes, play critical roles in bacterial growth and environmental adaptation. However, the underlying mechanism by which such regulators interact with their cognate DNA remains to be elucidated. In this study, we solved the crystal structure of a broad regulator Ms6564 in Mycobacterium smegmatis and its protein-operator complex at resolutions of 1.9 and 2.5 Å, respectively. Similar to other typical TetR family regulators, two dimeric Ms6564 molecules were found to bind to opposite sides of target DNA. However, the recognition helix of Ms6564 inserted only slightly into the DNA major groove. Unexpectedly, 11 disordered water molecules bridged the interface of TetR family regulator DNA. Although the DNA was deformed upon Ms6564 binding, it still retained the conformation of B-form DNA. Within the DNA-binding domain of Ms6564, only two amino acids residues directly interacted with the bases of cognate DNA. Lys-47 was found to be essential for the specific DNA binding ability of Ms6564. These data indicate that Ms6564 can bind DNA with strong affinity but makes flexible contacts with DNA. Our study suggests that Ms6564 might slide more easily along the genomic DNA and extensively regulate the expression of diverse genes in M. smegmatis.
- National Key Laboratory of Agricultural Microbiology, Center for Proteomics Research, College of Life Science and Technology, Huazhong Agricultural University, Wuhan 430070, China.
Organizational Affiliation: