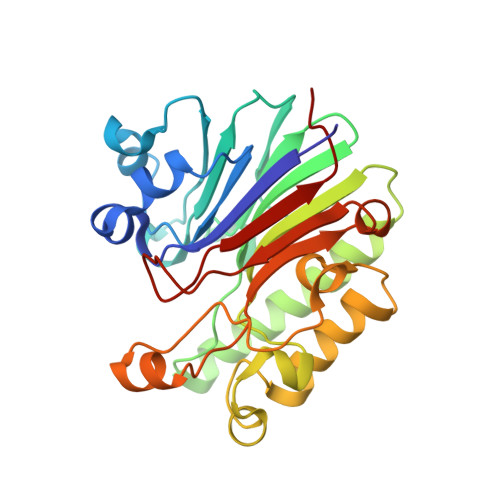
The Pseudomonas aeruginosa Catabolite Repression Control Protein Crc Is Devoid of RNA Binding Activity
Milojevic, T., Grishkovskaya, I., Sonnleitner, E., Djinovic-Carugo, K., Blasi, U.(2013) PLoS One 8: e64609-e64609
- PubMed: 23717639
- DOI: https://doi.org/10.1371/journal.pone.0064609
- Primary Citation of Related Structures:
4JG3 - PubMed Abstract:
The Crc protein has been shown to mediate catabolite repression control in Pseudomonas, leading to a preferential assimilation of carbon sources. It has been suggested that Crc acts as a translational repressor of mRNAs, encoding functions involved in uptake and breakdown of different carbon sources. Moreover, the regulatory RNA CrcZ, the level of which is increased in the presence of less preferred carbon sources, was suggested to bind to and sequester Crc, resulting in a relief of catabolite repression. Here, we determined the crystal structure of Pseudomonas aeruginosa Crc, a member of apurinic/apyrimidinic (AP) endonuclease family, at 1.8 Å. Although Crc displays high sequence similarity with its orthologs, there are amino acid alterations in the area corresponding to the active site in AP proteins. Unlike typical AP endonuclease family proteins, Crc has a reduced overall positive charge and the conserved positively charged amino-acid residues of the DNA-binding surface of AP proteins are partially substituted by negatively charged, polar and hydrophobic residues. Crc protein purified to homogeneity from P. aeruginosa did neither display DNase activity, nor did it bind to previously identified RNA substrates. Rather, the RNA chaperone Hfq was identified as a contaminant in His-tagged Crc preparations purified by one step Ni-affinity chromatography from Escherichia coli, and was shown to account for the RNA binding activity observed with the His-Crc preparations. Taken together, these data challenge a role of Crc as a direct translational repressor in carbon catabolite repression in P. aeruginosa.
- Department of Microbiology, Immunobiology and Genetics, Max F Perutz Laboratories, University of Vienna, Vienna, Austria.
Organizational Affiliation: