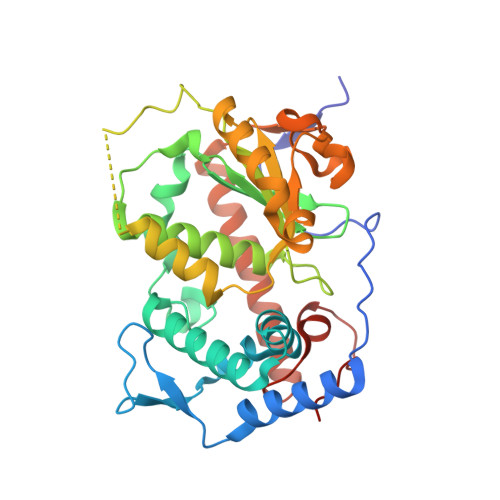
Structural determinants for phosphatidylinositol recognition by Sfh3 and substrate-induced dimer-monomer transition during lipid transfer cycles.
Yang, H., Tong, J., Leonard, T.A., Im, Y.J.(2013) FEBS Lett 587: 1610-1616
- PubMed: 23603387
- DOI: https://doi.org/10.1016/j.febslet.2013.04.009
- Primary Citation Related Structures:
4J7P, 4J7Q - PubMed Abstract:
Sec14 family homologs are the major yeast phosphatidylinositol/phosphatidylcholine transfer proteins regulating lipid metabolism and vesicle trafficking. The structure of Saccharomyces cerevisiae Sfh3 displays a conserved Sec14 scaffold and reveals determinants for the specific recognition of phosphatidylinositol ligand. Apo-Sfh3 forms a dimer through the hydrophobic interaction of gating helices. Binding of phosphatidylinositol leads to dissociation of the dimer into monomers in a reversible manner. This study suggests that the substrate induced dimer-monomer transformation is an essential part of lipid transfer cycles by Sfh3.
- College of Pharmacy, Chonnam National University, Gwangju 500-757, South Korea.
Organizational Affiliation: