Structural basis of protein phosphatase 2A stable latency.
Jiang, L., Stanevich, V., Satyshur, K.A., Kong, M., Watkins, G.R., Wadzinski, B.E., Sengupta, R., Xing, Y.(2013) Nat Commun 4: 1699-1699
- PubMed: 23591866 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms2663
- Primary Citation Related Structures:
4IYP - PubMed Abstract:
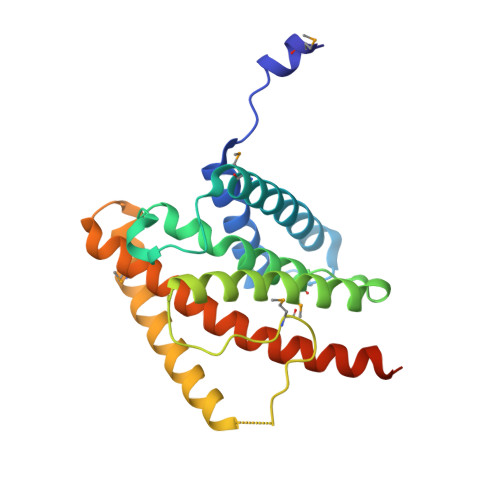
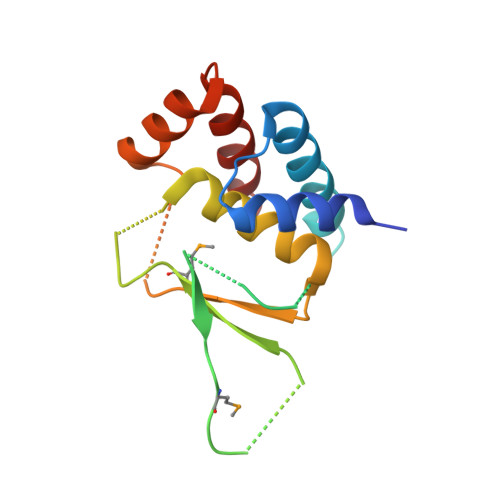
The catalytic subunit of protein phosphatase 2A (PP2Ac) is stabilized in a latent form by α4, a regulatory protein essential for cell survival and biogenesis of all PP2A complexes. Here we report the structure of α4 bound to the N-terminal fragment of PP2Ac. This structure suggests that α4 binding to the full-length PP2Ac requires local unfolding near the active site, which perturbs the scaffold subunit binding site at the opposite surface via allosteric relay. These changes stabilize an inactive conformation of PP2Ac and convert oligomeric PP2A complexes to the α4 complex upon perturbation of the active site. The PP2Ac-α4 interface is essential for cell survival and sterically hinders a PP2A ubiquitination site, important for the stability of cellular PP2Ac. Our results show that α4 is a scavenger chaperone that binds to and stabilizes partially folded PP2Ac for stable latency, and reveal a mechanism by which α4 regulates cell survival, and biogenesis and surveillance of PP2A holoenzymes.
- McArdle Laboratory, Department of Oncology, University of Wisconsin-Madison, School of Medicine and Public Health, Madison, Wisconsin 53706, USA.
Organizational Affiliation: