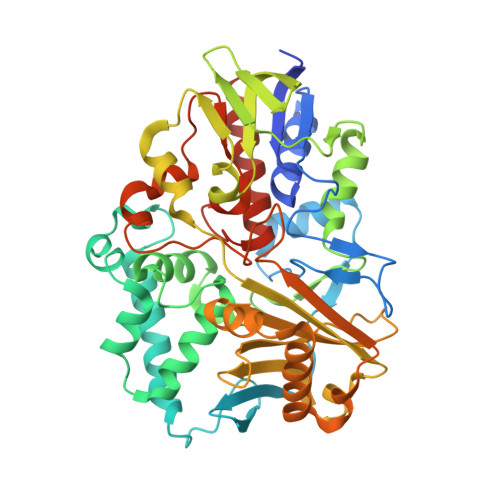
Structural Analysis of a Novel Cyclohexylamine Oxidase from Brevibacterium oxydans IH-35A.
Mirza, I.A., Burk, D.L., Xiong, B., Iwaki, H., Hasegawa, Y., Grosse, S., Lau, P.C., Berghuis, A.M.(2013) PLoS One 8: e60072-e60072
- PubMed: 23555888
- DOI: https://doi.org/10.1371/journal.pone.0060072
- Primary Citation Related Structures:
4I58, 4I59 - PubMed Abstract:
Cyclohexylamine oxidase (CHAO) is a flavoprotein first described in Brevibacterium oxydans strain IH-35A that carries out the initial step of the degradation of the industrial chemical cyclohexylamine to cyclohexanone. We have cloned and expressed in Escherichia coli the CHAO-encoding gene (chaA) from B. oxydans, purified CHAO and determined the structures of both the holoenzyme form of the enzyme and a product complex with cyclohexanone. CHAO is a 50 kDa monomer with a PHBH fold topology. It belongs to the flavin monooxygenase family of enzymes and exhibits high substrate specificity for alicyclic amines and sec-alkylamines. The overall structure is similar to that of other members of the flavin monooxygenase family, but lacks either of the C- or N-terminal extensions observed in these enzymes. Active site features of the flavin monooxygenase family are conserved in CHAO, including the characteristic aromatic cage. Differences in the orientations of residues of the CHAO aromatic cage result in a substrate-binding site that is more open than those of its structural relatives. Since CHAO has a buried hydrophobic active site with no obvious route for substrates and products, a random acceleration molecular dynamics simulation has been used to identify a potential egress route. The path identified includes an intermediate cavity and requires transient conformation changes in a shielding loop and a residue at the border of the substrate-binding cavity. These results provide a foundation for further studies with CHAO aimed at identifying features determining substrate specificity and for developing the biocatalytic potential of this enzyme.
- Department of Biochemistry and Groupe de Recherche Axé sur la Structure des Protéines, McGill University, Montreal, Quebec, Canada.
Organizational Affiliation: