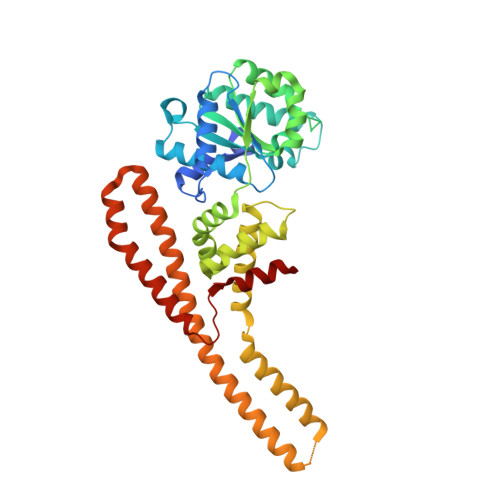
The molecular mechanism of hsp100 chaperone inhibition by the prion curing agent guanidinium chloride.
Zeymer, C., Werbeck, N.D., Schlichting, I., Reinstein, J.(2013) J Biological Chem 288: 7065-7076
- PubMed: 23341453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.432583
- Primary Citation Related Structures:
4HSE - PubMed Abstract:
The Hsp100 chaperones ClpB and Hsp104 utilize the energy from ATP hydrolysis to reactivate aggregated proteins in concert with the DnaK/Hsp70 chaperone system, thereby playing an important role in protein quality control. They belong to the family of AAA+ proteins (ATPases associated with various cellular activities), possess two nucleotide binding domains per monomer (NBD1 and NBD2), and oligomerize into hexameric ring complexes. Furthermore, Hsp104 is involved in yeast prion propagation and inheritance. It is well established that low concentrations of guanidinium chloride (GdmCl) inhibit the ATPase activity of Hsp104, leading to so called "prion curing," the loss of prion-related phenotypes. Here, we present mechanistic details about the Hsp100 chaperone inhibition by GdmCl using the Hsp104 homolog ClpB from Thermus thermophilus. Initially, we demonstrate that NBD1 of ClpB, which was previously considered inactive as a separately expressed construct, is a fully active ATPase on its own. Next, we show that only NBD1, but not NBD2, is affected by GdmCl. We present a crystal structure of ClpB NBD1 in complex with GdmCl and ADP, showing that the Gdm(+) ion binds specifically to the active site of NBD1. A conserved essential glutamate residue is involved in this interaction. Additionally, Gdm(+) interacts directly with the nucleotide, thereby increasing the nucleotide binding affinity of NBD1. We propose that both the interference with the essential glutamate and the modulation of nucleotide binding properties in NBD1 is responsible for the GdmCl-specific inhibition of Hsp100 chaperones.
- Department of Biomolecular Mechanisms, Max Planck Institute for Medical Research, 69120 Heidelberg, Germany.
Organizational Affiliation: