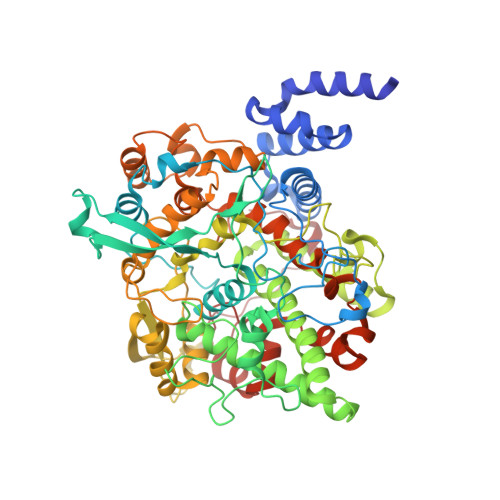
The crystal structure of alpha-Dioxygenase provides insight into diversity in the cyclooxygenase-peroxidase superfamily.
Goulah, C.C., Zhu, G., Koszelak-Rosenblum, M., Malkowski, M.G.(2013) Biochemistry 52: 1364-1372
- PubMed: 23373518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi400013k
- Primary Citation Related Structures:
4HHR, 4HHS - PubMed Abstract:
α-Dioxygenases (α-DOX) oxygenate fatty acids into 2(R)-hydroperoxides. Despite the low level of sequence identity, α-DOX share common catalytic features with cyclooxygenases (COX), including the use of a tyrosyl radical during catalysis. We determined the X-ray crystal structure of Arabidopsis thaliana α-DOX to 1.5 Å resolution. The α-DOX structure is monomeric, predominantly α-helical, and comprised of two domains. The base domain exhibits a low degree of structural homology with the membrane-binding domain of COX but lies in a similar position with respect to the catalytic domain. The catalytic domain shows the highest degree of similarity with the COX catalytic domain, where 21 of the 22 α-helical elements are conserved. Helices H2, H6, H8, and H17 form the heme binding cleft and walls of the active site channel. His-318, Thr-323, and Arg-566 are located near the catalytic tyrosine, Tyr-386, at the apex of the channel, where they interact with a chloride ion. Substitutions at these positions coupled with kinetic analyses confirm previous hypotheses that implicate these residues as being involved in binding and orienting the carboxylate group of the fatty acid for optimal catalysis. Unique to α-DOX is the presence of two extended inserts on the surface of the enzyme that restrict access to the distal face of the heme, providing an explanation for the observed reduced peroxidase activity of the enzyme. The α-DOX structure represents the first member of the α-DOX subfamily to be structurally characterized within the cyclooxygenase-peroxidase family of heme-containing proteins.
- Hauptman-Woodward Medical Research Institute, The State University of New York at Buffalo, New York 14203, United States.
Organizational Affiliation: