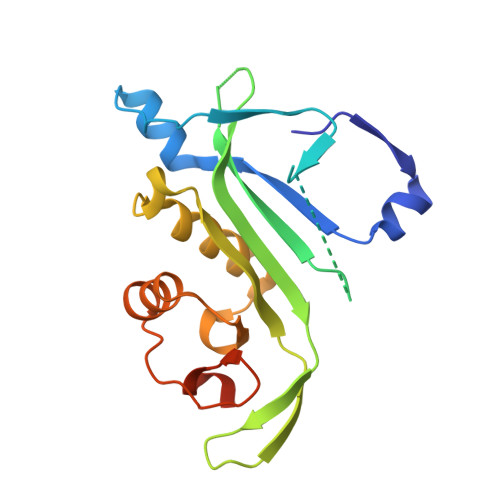
Crystal structures of tubulin acetyltransferase reveal a conserved catalytic core and the plasticity of the essential N terminus.
Kormendi, V., Szyk, A., Piszczek, G., Roll-Mecak, A.(2012) J Biological Chem 287: 41569-41575
- PubMed: 23105108 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C112.421222
- Primary Citation Related Structures:
4H6U, 4H6Z - PubMed Abstract:
Tubulin acetyltransferase (TAT) acetylates Lys-40 of α-tubulin in the microtubule lumen. TAT is inefficient, and its activity is enhanced when tubulin is incorporated in microtubules. Acetylation is associated with stable microtubules and regulates the binding of microtubule motors and associated proteins. TAT is important in neuronal polarity and mechanosensation, and decreased tubulin acetylation levels are associated with axonal transport defects and neurodegeneration. We present the first structure of TAT in complex with acetyl-CoA (Ac-CoA) at 2.7 Å resolution. The structure reveals a conserved stable catalytic core shared with other GCN5 superfamily acetyltransferases consisting of a central β-sheet flanked by α-helices and a C-terminal β-hairpin unique to TAT. Structure-guided mutagenesis establishes the molecular determinants for Ac-CoA and tubulin substrate recognition. The wild-type TAT construct is a monomer in solution. We identify a metastable interface between the conserved core and N-terminal domain that modulates the oligomerization of TAT in solution and is essential for activity. The 2.45 Å resolution structure of an inactive TAT construct with an active site point mutation near this interface reveals a domain-swapped dimer in which the functionally essential N terminus shows evidence of marked structural plasticity. The sequence segment corresponding to this structurally plastic region in TAT has been implicated in substrate recognition in other GCN5 superfamily acetyltransferases. Our structures provide a rational platform for the mechanistic dissection of TAT activity and the design of TAT inhibitors with therapeutic potential in neuronal regeneration.
- Cell Biology and Biophysics Unit, NINDS, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: