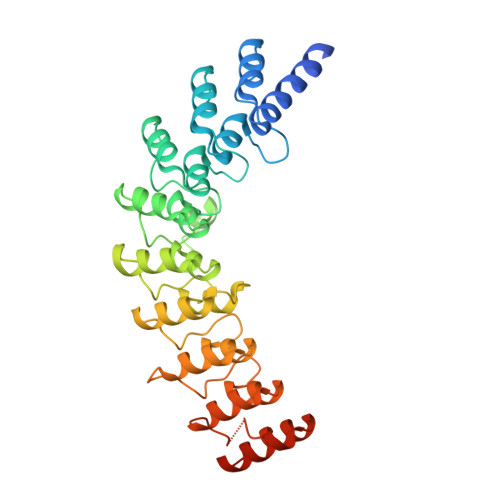
Innate Immune Messenger 2-5A Tethers Human RNase L into Active High-Order Complexes.
Han, Y., Whitney, G., Donovan, J., Korennykh, A.(2012) Cell Rep 2: 902-913
- PubMed: 23084743
- DOI: https://doi.org/10.1016/j.celrep.2012.09.004
- Primary Citation of Related Structures:
4G8K, 4G8L - PubMed Abstract:
2',5'-linked oligoadenylates (2-5As) serve as conserved messengers of pathogen presence in the mammalian innate immune system. 2-5As induce self-association and activation of RNase L, which cleaves cytosolic RNA and promotes the production of interferons (IFNs) and cytokines driven by the transcription factors IRF-3 and NF-κB. We report that human RNase L is activated by forming high-order complexes, reminiscent of the mode of activation of the phylogenetically related transmembrane kinase/RNase Ire1 in the unfolded protein response. We describe crystal structures determined at 2.4 Å and 2.8 Å resolution, which show that two molecules of 2-5A at a time tether RNase L monomers via the ankyrin-repeat (ANK) domain. Each ANK domain harbors two distinct sites for 2-5A recognition that reside 50 Å apart. These data reveal a function for the ANK domain as a 2-5A-sensing homo-oligomerization device and describe a nonlinear, ultrasensitive regulation in the 2-5A/RNase L system poised for amplification of the IFN response.
- Department of Molecular Biology, Princeton University, 216 Schultz Laboratory, Princeton, NJ 08540, USA.
Organizational Affiliation: