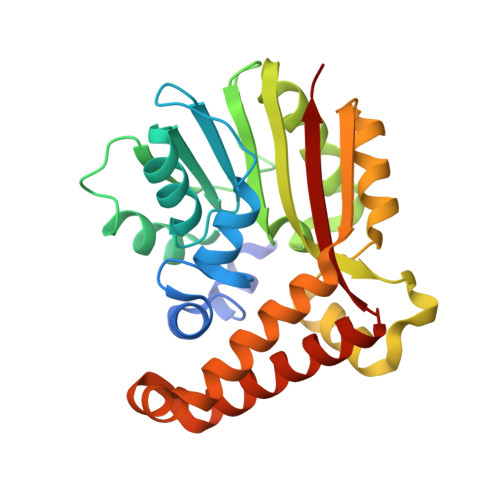
Crystal structure of phosphoethanolamine methyltransferase from Plasmodium falciparum in complex with amodiaquine.
Lee, S.G., Alpert, T.D., Jez, J.M.(2012) Bioorg Med Chem Lett 22: 4990-4993
- PubMed: 22771008 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2012.06.032
- Primary Citation Related Structures:
4FGZ - PubMed Abstract:
Phosphoethanolamine N-methyltransferase (PMT) is essential for phospholipid biogenesis in the malarial parasite Plasmodium falciparum. PfPMT catalyzes the triple methylation of phosphoethanolamine to produce phosphocholine, which is then used for phosphatidylcholine synthesis. Here we describe the 2.0Å resolution X-ray crystal structure of PfPMT in complex with amodiaquine. To better characterize inhibition of PfPMT by amodiaquine, we determined the IC(50) values of a series of aminoquinolines using a direct radiochemical assay. Both structural and functional analyses provide a possible approach for the development of new small molecule inhibitors of PfPMT.
- Department of Biology, Washington University, St. Louis, MO 63130, USA.
Organizational Affiliation: