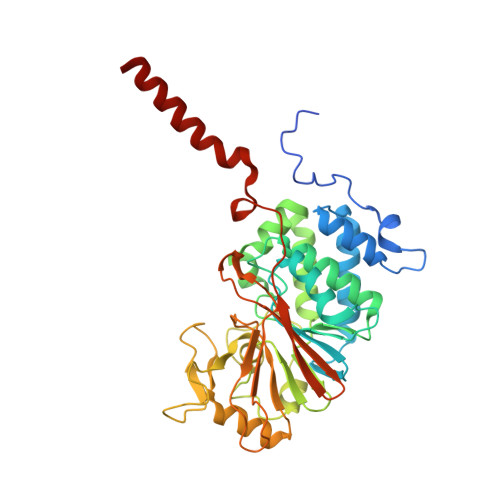
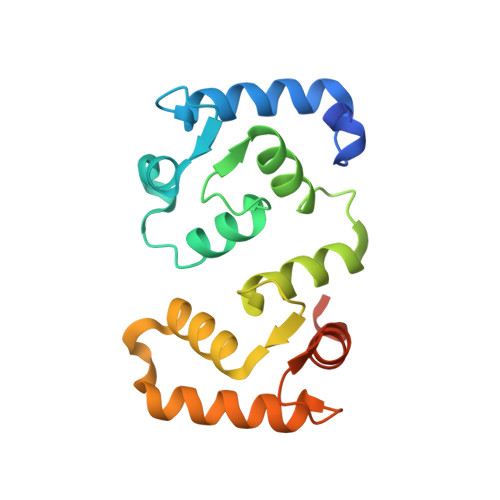
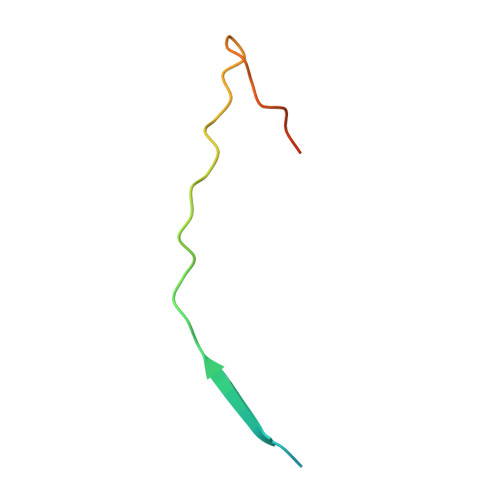
The molecular mechanism of substrate engagement and immunosuppressant inhibition of calcineurin.
Grigoriu, S., Bond, R., Cossio, P., Chen, J.A., Ly, N., Hummer, G., Page, R., Cyert, M.S., Peti, W.(2013) PLoS Biol 11: e1001492-e1001492
- PubMed: 23468591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1001492
- Primary Citation Related Structures:
4F0Z - PubMed Abstract:
Ser/thr phosphatases dephosphorylate their targets with high specificity, yet the structural and sequence determinants of phosphosite recognition are poorly understood. Calcineurin (CN) is a conserved Ca(2+)/calmodulin-dependent ser/thr phosphatase and the target of immunosuppressants, FK506 and cyclosporin A (CSA). To investigate CN substrate recognition we used X-ray crystallography, biochemistry, modeling, and in vivo experiments to study A238L, a viral protein inhibitor of CN. We show that A238L competitively inhibits CN by occupying a critical substrate recognition site, while leaving the catalytic center fully accessible. Critically, the 1.7 Å structure of the A238L-CN complex reveals how CN recognizes residues in A238L that are analogous to a substrate motif, "LxVP." The structure enabled modeling of a peptide substrate bound to CN, which predicts substrate interactions beyond the catalytic center. Finally, this study establishes that "LxVP" sequences and immunosuppressants bind to the identical site on CN. Thus, FK506, CSA, and A238L all prevent "LxVP"-mediated substrate recognition by CN, highlighting the importance of this interaction for substrate dephosphorylation. Collectively, this work presents the first integrated structural model for substrate selection and dephosphorylation by CN and lays the groundwork for structure-based development of new CN inhibitors.
- Department of Molecular Pharmacology, Physiology and Biotechnology, Brown University, Providence, Rhode Island, USA.
Organizational Affiliation: