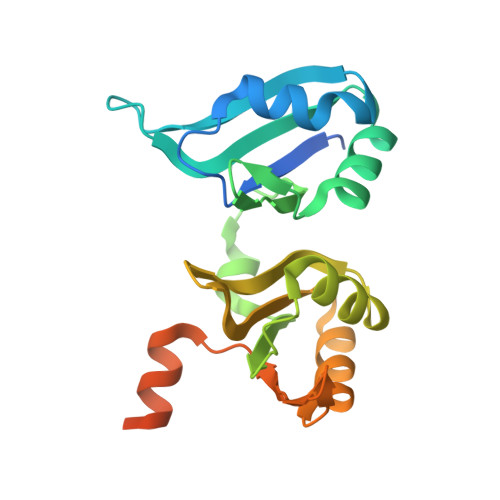
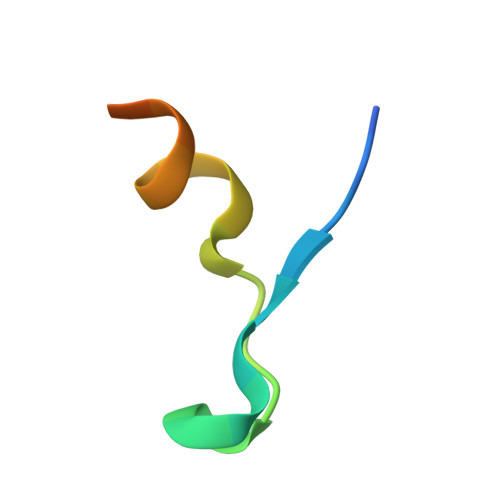
Interdomain Allostery Promotes Assembly of the Poly(A) mRNA Complex with PABP and eIF4G.
Safaee, N., Kozlov, G., Noronha, A.M., Xie, J., Wilds, C.J., Gehring, K.(2012) Mol Cell 48: 375-386
- PubMed: 23041282 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2012.09.001
- Primary Citation Related Structures:
4F02, 4F25, 4F26 - PubMed Abstract:
Many RNA-binding proteins contain multiple single-strand nucleic acid-binding domains and assemble into large multiprotein messenger ribonucleic acid protein (mRNP) complexes. The mechanisms underlying the self-assembly of these complexes are largely unknown. In eukaryotes, the association of the translation factors polyadenylate-binding protein-1 (PABP) and eIF4G is essential for high-level expression of polyadenylated mRNAs. Here, we report the crystal structure of the ternary complex poly(A)(11)·PABP(1-190)·eIF4G(178-203) at 2.0 Å resolution. Our NMR and crystallographic data show that eIF4G interacts with the RRM2 domain of PABP. Analysis of the interaction by small-angle X-ray scattering, isothermal titration calorimetry, and electromobility shift assays reveals that this interaction is allosterically regulated by poly(A) binding to PABP. Furthermore, we have confirmed the importance of poly(A) for the endogenous PABP and eIF4G interaction in immunoprecipitation experiments using HeLa cell extracts. Our findings reveal interdomain allostery as a mechanism for cooperative assembly of RNP complexes.
- Department of Biochemistry, McGill University, Montréal, QC H3G 0B1, Canada.
Organizational Affiliation: