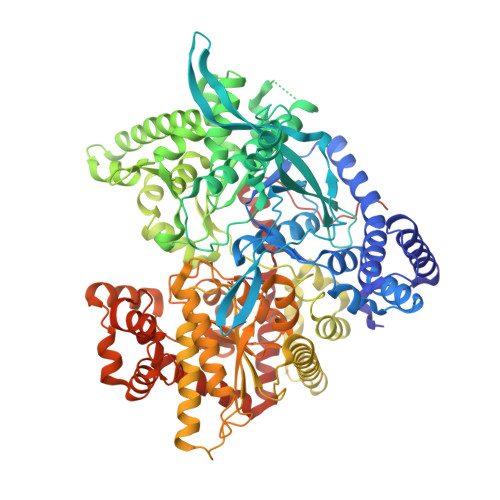
The binding of C5-alkynyl and alkylfurano[2,3-d]pyrimidine glucopyranonucleosides to glycogen phosphorylase b: Synthesis, biochemical and biological assessment.
Kantsadi, A.L., Manta, S., Psarra, A.M., Dimopoulou, A., Kiritsis, C., Parmenopoulou, V., Skamnaki, V.T., Zoumpoulakis, P., Zographos, S.E., Leonidas, D.D., Komiotis, D.(2012) Eur J Med Chem 54: 740-749
- PubMed: 22770609
- DOI: https://doi.org/10.1016/j.ejmech.2012.06.029
- Primary Citation of Related Structures:
4EJ2, 4EKE, 4EKY, 4EL0, 4EL5 - PubMed Abstract:
C5-alkynyl and alkylfurano[2,3-d]pyrimidine glucopyranonucleosides have been synthesized and studied as inhibitors of glycogen phosphorylase b (GPb). Kinetic experiments have shown that most of these compounds were low micromolar inhibitors of the enzyme. The best inhibitor was 1-(β-D-glucopyranosyl)-5-ethynyluracil (K(i)=4.7 μM). Crystallographic analysis of these compounds in complex with GPb revealed that inhibitors with a long C5-alkynyl group exploited interactions with β-pocket of the active site and induced significant conformational changes of the 280s loop compared to GPb in complex with compounds with a short C5-alkynyl group. The results highlight the importance in the length of the aliphatic groups used to enhance inhibitory potency for the exploitation of the hydrophobic β-pocket. The best of the inhibitors had also a moderate effect on glycogenolysis in the cellular lever with an IC(50) value of 291.4 μM.
- Department of Biochemistry and Biotechnology, University of Thessaly, 26 Ploutonos Str. 41221 Larissa, Greece.
Organizational Affiliation: