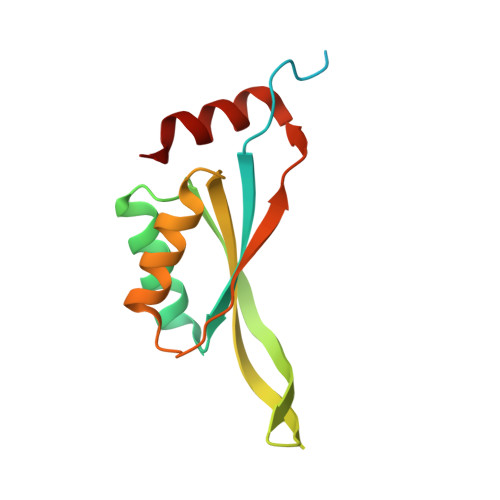
Structure of a CutA1 divalent-cation tolerance protein from Cryptosporidium parvum, the protozoal parasite responsible for cryptosporidiosis.
Buchko, G.W., Abendroth, J., Clifton, M.C., Robinson, H., Zhang, Y., Hewitt, S.N., Staker, B.L., Edwards, T.E., Van Voorhis, W.C., Myler, P.J.(2015) Acta Crystallogr Sect F Struct Biol Cryst Commun 71: 522-530
- PubMed: 25945704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14028210
- Primary Citation Related Structures:
4E98 - PubMed Abstract:
Cryptosporidiosis is an infectious disease caused by protozoan parasites of the Cryptosporidium genus. Infection is associated with mild to severe diarrhea that usually resolves spontaneously in healthy human adults, but may lead to severe complications in young children and in immunocompromised patients. The genome of C. parvum contains a gene, CUTA_CRYPI, that may play a role in regulating the intracellular concentration of copper, which is a toxic element in excess. Here, the crystal structure of this CutA1 protein, Cp-CutA1, is reported at 2.0 Å resolution. As observed for other CutA1 structures, the 117-residue protein is a trimer with a core ferrodoxin-like fold. Circular dichroism spectroscopy shows little, in any, unfolding of Cp-CutA1 up to 353 K. This robustness is corroborated by (1)H-(15)N HSQC spectra at 333 K, which are characteristic of a folded protein, suggesting that NMR spectroscopy may be a useful tool to further probe the function of the CutA1 proteins. While robust, Cp-CutA1 is not as stable as the homologous protein from a hyperthermophile, perhaps owing to a wide β-bulge in β2 that protrudes Pro48 and Ser49 outside the β-sheet.
- Seattle Structural Genomics Center for Infectious Disease, USA.
Organizational Affiliation: