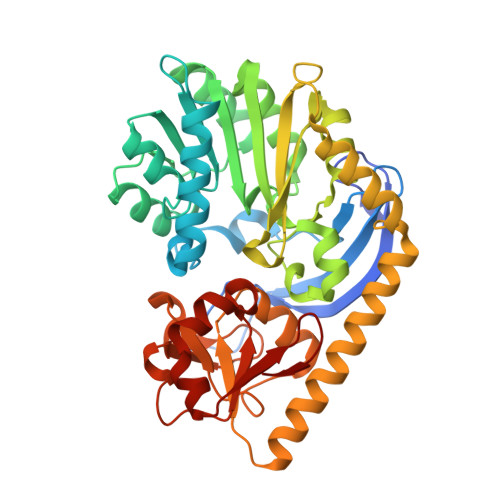
Probing the catalytic mechanism of a C-3'-methyltransferase involved in the biosynthesis of D-tetronitrose.
Bruender, N.A., Holden, H.M.(2012) Protein Sci 21: 876-886
- PubMed: 22495991
- DOI: https://doi.org/10.1002/pro.2074
- Primary Citation of Related Structures:
4E2W, 4E2X, 4E2Y, 4E2Z, 4E30, 4E31, 4E32, 4E33 - PubMed Abstract:
D-Tetronitrose is a nitro-containing tetradeoxysugar found attached to the antitumor and antibacterial agent tetrocarcin A. The biosynthesis of this highly unusual sugar in Micromonospora chalcea requires 10 enzymes. The fifth step in the pathway involves the transfer of a methyl group from S-adenosyl-L-methionine (SAM) to the C-3' carbon of dTDP-3-amino-2,3,6-trideoxy-4-keto-D-glucose. The enzyme responsible for this transformation is referred to as TcaB9. It is a monomeric enzyme with a molecular architecture based around three domains. The N-terminal motif contains a binding site for a structural zinc ion. The middle- and C-terminal domains serve to anchor the SAM and dTDP-sugar ligands, respectively, to the protein, and the active site of TcaB9 is wedged between these two regions. For this investigation, the roles of Tyr 76, His 181, Tyr 222, Glu 224, and His 225, which form the active site of TcaB9, were probed by site-directed mutagenesis, kinetic analyses, and X-ray structural studies. In addition, two ternary complexes of the enzyme with bound S-adenosyl-L-homocysteine and either dTDP-3-amino-2,3,6-trideoxy-4-keto-D-glucose or dTDP-3-amino-2,3,6-trideoxy-D-galactose were determined to 1.5 or 1.6 Å resolution, respectively. Taken together, these investigations highlight the important role of His 225 in methyl transfer. In addition, the structural data suggest that the methylation reaction occurs via retention of configuration about the C-3' carbon of the sugar.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: