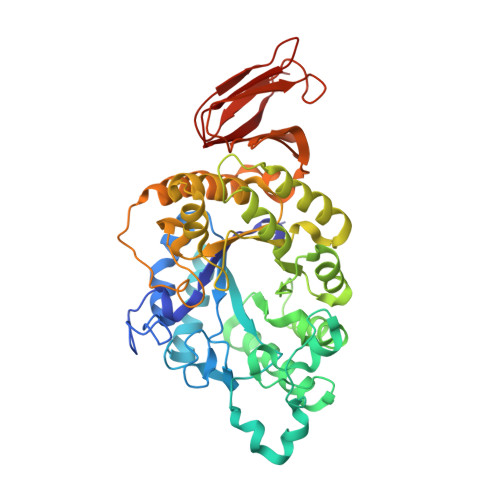
Crystal structure of a compact alpha-amylase from Geobacillus thermoleovorans.
Mok, S.C., Teh, A.H., Saito, J.A., Najimudin, N., Alam, M.(2013) Enzyme Microb Technol 53: 46-54
- PubMed: 23683704 Search on PubMed
- DOI: https://doi.org/10.1016/j.enzmictec.2013.03.009
- Primary Citation Related Structures:
4E2O - PubMed Abstract:
A truncated form of an α-amylase, GTA, from thermophilic Geobacillus thermoleovorans CCB_US3_UF5 was biochemically and structurally characterized. The recombinant GTA, which lacked both the N- and C-terminal transmembrane regions, functioned optimally at 70°C and pH 6.0. While enzyme activity was not enhanced by the addition of CaCl2, GTA's thermostability was significantly improved in the presence of CaCl2. The structure, in complex with an acarbose-derived pseudo-hexasaccharide, consists of the typical three domains and binds one Ca(2+) ion. This Ca(2+) ion was strongly bound and not chelated by EDTA. A predicted second Ca(2+)-binding site, however, was disordered. With limited subsites, two novel substrate-binding residues, Y147 and Y182, may help increase substrate affinity. No distinct starch-binding domain is present, although two regions rich in aromatic residues have been observed. GTA, with a smaller domain B and several shorter loops compared to other α-amylases, has one of the most compact α-amylase folds that may contribute greatly to its tight Ca(2+) binding and thermostability.
- Centre for Chemical Biology, Universiti Sains Malaysia, 11800 Penang, Malaysia.
Organizational Affiliation: