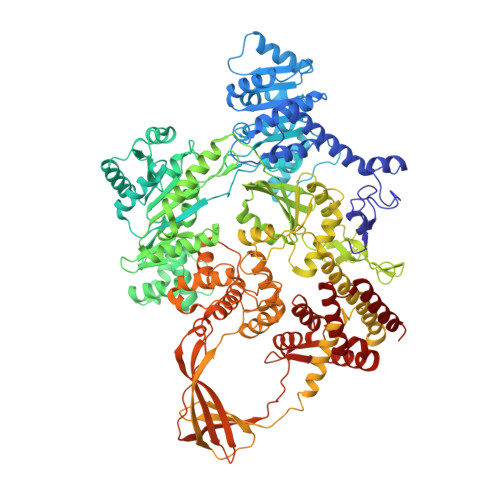
Crystal structures of Thermotoga maritima reverse gyrase: inferences for the mechanism of positive DNA supercoiling.
Rudolph, M.G., Del Toro Duany, Y., Jungblut, S.P., Ganguly, A., Klostermeier, D.(2013) Nucleic Acids Res 41: 1058-1070
- PubMed: 23209025
- DOI: https://doi.org/10.1093/nar/gks1073
- Primary Citation Related Structures:
4DDT, 4DDU, 4DDV, 4DDW, 4DDX - PubMed Abstract:
Reverse gyrase is an ATP-dependent topoisomerase that is unique to hyperthermophilic archaea and eubacteria. The only reverse gyrase structure determined to date has revealed the arrangement of the N-terminal helicase domain and the C-terminal topoisomerase domain that intimately cooperate to generate the unique function of positive DNA supercoiling. Although the structure has elicited hypotheses as to how supercoiling may be achieved, it lacks structural elements important for supercoiling and the molecular mechanism of positive supercoiling is still not clear. We present five structures of authentic Thermotoga maritima reverse gyrase that reveal a first view of two interacting zinc fingers that are crucial for positive DNA supercoiling. The so-called latch domain, which connects the helicase and the topoisomerase domains is required for their functional cooperation and presents a novel fold. Structural comparison defines mobile regions in parts of the helicase domain, including a helical insert and the latch that are likely important for DNA binding during catalysis. We show that the latch, the helical insert and the zinc fingers contribute to the binding of DNA to reverse gyrase and are uniquely placed within the reverse gyrase structure to bind and guide DNA during strand passage. A possible mechanism for positive supercoiling by reverse gyrases is presented.
- pRED, Pharma Research and Early Development, Discovery Technologies, Grenzacherstrasse 124, CH-4070 Basel, Switzerland.
Organizational Affiliation: