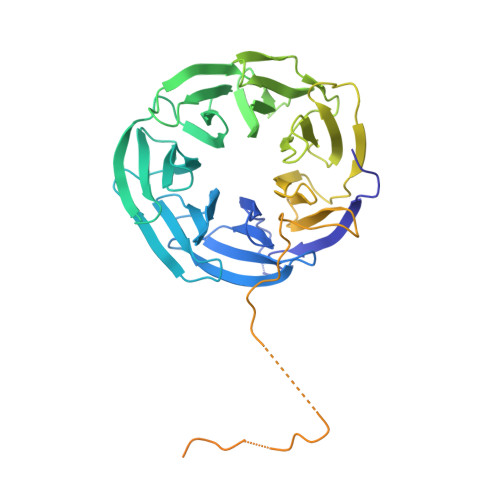
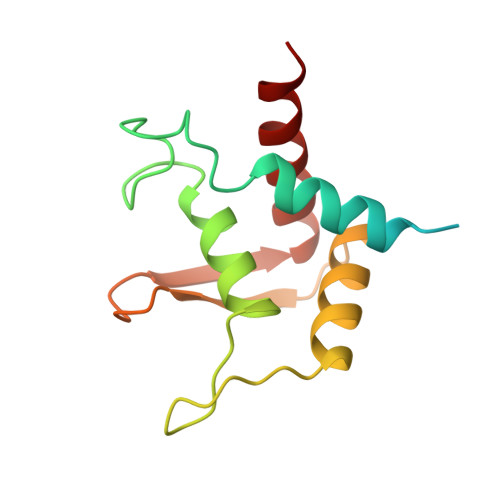
An Asymmetric Pan3 Dimer Recruits a Single Pan2 Exonuclease to Mediate Mrna Deadenylation and Decay.
Jonas, S., Christie, M., Peter, D., Bhandari, D., Loh, B., Huntzinger, E., Weichenrieder, O., Izaurralde, E.(2014) Nat Struct Mol Biol 21: 599
- PubMed: 24880343 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2837
- Primary Citation Related Structures:
4CZV, 4CZW, 4CZX, 4CZY, 4D0K - PubMed Abstract:
The PAN2-PAN3 complex functions in general and microRNA-mediated mRNA deadenylation. However, mechanistic insight into PAN2 and its complex with the asymmetric PAN3 dimer is lacking. Here, we describe crystal structures that show that Neurospora crassa PAN2 comprises two independent structural units: a C-terminal catalytic unit and an N-terminal assembly unit that engages in a bipartite interaction with PAN3 dimers. The catalytic unit contains the exonuclease domain in an intimate complex with a potentially modulatory ubiquitin-protease-like domain. The assembly unit contains a WD40 propeller connected to an adaptable linker. The propeller contacts the PAN3 C-terminal domain, whereas the linker reinforces the asymmetry of the PAN3 dimer and prevents the recruitment of a second PAN2 molecule. Functional data indicate an essential role for PAN3 in coordinating PAN2-mediated deadenylation with subsequent steps in mRNA decay, which lead to complete mRNA degradation.
- 1] Department of Biochemistry, Max Planck Institute for Developmental Biology, Tübingen, Germany. [2].
Organizational Affiliation: