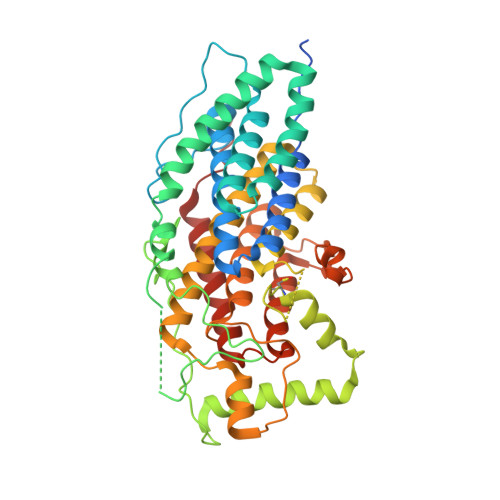
The Bacterial Vanadium Iodoperoxidase from the Marine Flavobacteriaceae Zobellia Galactanivorans Reveals Novel Molecular and Evolutionary Features of Halide Specificity in This Enzyme Family.
Fournier, J.B., Rebuffet, E., Delage, L., Grijol, R., Meslet-Cladiere, L., Rzonca, J., Potin, P., Michel, G., Czjzek, M., Leblanc, C.(2014) Appl Environ Microbiol 80: 7561
- PubMed: 25261522 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AEM.02430-14
- Primary Citation Related Structures:
4CIT, 4USZ - PubMed Abstract:
Vanadium haloperoxidases (VHPO) are key enzymes that oxidize halides and are involved in the biosynthesis of organo-halogens. Until now, only chloroperoxidases (VCPO) and bromoperoxidases (VBPO) have been characterized structurally, mainly from eukaryotic species. Three putative VHPO genes were predicted in the genome of the flavobacterium Zobellia galactanivorans, a marine bacterium associated with macroalgae. In a phylogenetic analysis, these putative bacterial VHPO were closely related to other VHPO from diverse bacterial phyla but clustered independently from eukaryotic algal VBPO and fungal VCPO. Two of these bacterial VHPO, heterogeneously produced in Escherichia coli, were found to be strictly specific for iodide oxidation. The crystal structure of one of these vanadium-dependent iodoperoxidases, Zg-VIPO1, was solved by multiwavelength anomalous diffraction at 1.8 Å, revealing a monomeric structure mainly folded into α-helices. This three-dimensional structure is relatively similar to those of VCPO of the fungus Curvularia inaequalis and of Streptomyces sp. and is superimposable onto the dimeric structure of algal VBPO. Surprisingly, the vanadate binding site of Zg-VIPO1 is strictly conserved with the fungal VCPO active site. Using site-directed mutagenesis, we showed that specific amino acids and the associated hydrogen bonding network around the vanadate center are essential for the catalytic properties and also the iodide specificity of Zg-VIPO1. Altogether, phylogeny and structure-function data support the finding that iodoperoxidase activities evolved independently in bacterial and algal lineages, and this sheds light on the evolution of the VHPO enzyme family.
- Sorbonne Universités, UPMC Université Paris 06, UMR 8227, Integrative Biology of Marine Models, Station Biologique, Roscoff, France; CNRS, UMR 8227, Integrative Biology of Marine Models, Station Biologique, Roscoff, France.
Organizational Affiliation: