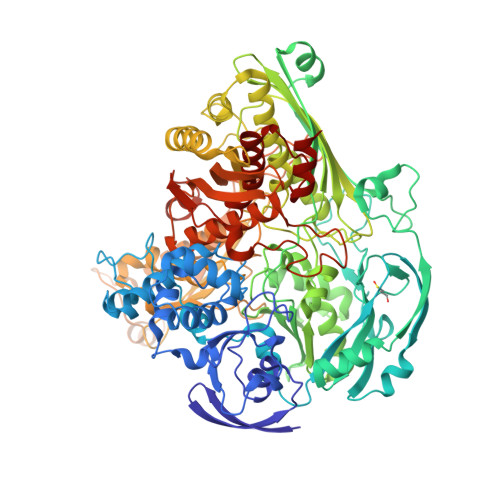
Kinetic and Structural Studies of Aldehyde Oxidoreductase from Desulfovibrio Gigas Reveal a Dithiolene-Based Chemistry for Enzyme Activation and Inhibition by H2O2.
Marangon, J., Correia, H.D., Brondino, C.D., Moura, J.J.G., Romao, M.J., Gonzalez, P.J., Santos-Silva, T.(2013) PLoS One 8: 83234
- PubMed: 24391748 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0083234
- Primary Citation Related Structures:
4C7Y, 4C7Z, 4C80 - PubMed Abstract:
Mononuclear Mo-containing enzymes of the xanthine oxidase (XO) family catalyze the oxidative hydroxylation of aldehydes and heterocyclic compounds. The molybdenum active site shows a distorted square-pyramidal geometry in which two ligands, a hydroxyl/water molecule (the catalytic labile site) and a sulfido ligand, have been shown to be essential for catalysis. The XO family member aldehyde oxidoreductase from Desulfovibrio gigas (DgAOR) is an exception as presents in its catalytically competent form an equatorial oxo ligand instead of the sulfido ligand. Despite this structural difference, inactive samples of DgAOR can be activated upon incubation with dithionite plus sulfide, a procedure similar to that used for activation of desulfo-XO. The fact that DgAOR does not need a sulfido ligand for catalysis indicates that the process leading to the activation of inactive DgAOR samples is different to that of desulfo-XO. We now report a combined kinetic and X-ray crystallographic study to unveil the enzyme modification responsible for the inactivation and the chemistry that occurs at the Mo site when DgAOR is activated. In contrast to XO, which is activated by resulfuration of the Mo site, DgAOR activation/inactivation is governed by the oxidation state of the dithiolene moiety of the pyranopterin cofactor, which demonstrates the non-innocent behavior of the pyranopterin in enzyme activity. We also showed that DgAOR incubation with dithionite plus sulfide in the presence of dioxygen produces hydrogen peroxide not associated with the enzyme activation. The peroxide molecule coordinates to molybdenum in a η(2) fashion inhibiting the enzyme activity.
- REQUIMTE/CQFB, Departamento de Química, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, Caparica, Setubal, Portugal.
Organizational Affiliation: