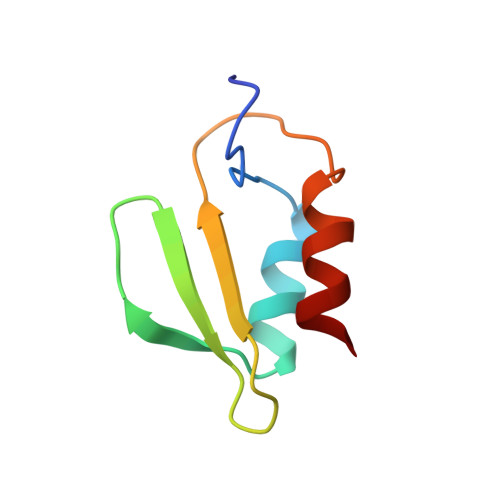
The Hica Toxin from Burkholderia Pseudomallei Has a Role in Persister Cell Formation.
Butt, A., Higman, V.A., Williams, C., Crump, M.P., Hemsley, C.M., Harmer, N., Titball, R.W.(2014) Biochem J 459: 333
- PubMed: 24502667 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20140073
- Primary Citation Related Structures:
4C26 - PubMed Abstract:
TA (toxin-antitoxin) systems are widely distributed amongst bacteria and are associated with the formation of antibiotic tolerant (persister) cells that may have involvement in chronic and recurrent disease. We show that overexpression of the Burkholderia pseudomallei HicA toxin causes growth arrest and increases the number of persister cells tolerant to ciprofloxacin or ceftazidime. Furthermore, our data show that persistence towards ciprofloxacin or ceftazidime can be differentially modulated depending on the level of induction of HicA expression. Deleting the hicAB locus from B. pseudomallei K96243 significantly reduced persister cell frequencies following exposure to ciprofloxacin, but not ceftazidime. The structure of HicA(H24A) was solved by NMR and forms a dsRBD-like (dsRNA-binding domain-like) fold, composed of a triple-stranded β-sheet, with two helices packed against one face. The surface of the protein is highly positively charged indicative of an RNA-binding protein and His24 and Gly22 were functionality important residues. This is the first study demonstrating a role for the HicAB system in bacterial persistence and the first structure of a HicA protein that has been experimentally characterized.
- *Biosciences, College of Life and Environmental Sciences, University of Exeter, Geoffrey Pope Building, Stocker Road, Exeter EX4 4QD, U.K.
Organizational Affiliation: