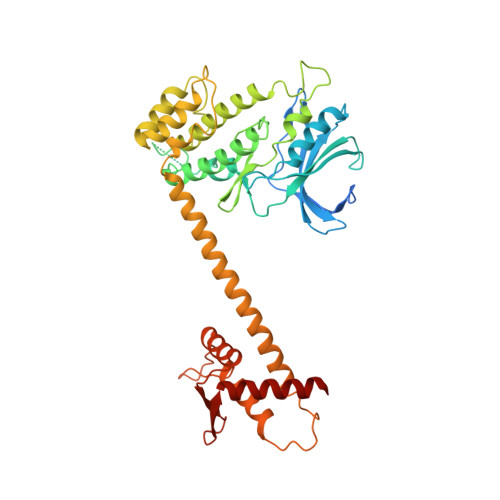
Structure of the Pan3 Pseudokinase Reveals the Basis for Interactions with the Pan2 Deadenylase and the Gw182 Proteins
Christie, M., Boland, A., Huntzinger, E., Weichenrieder, O., Izaurralde, E.(2013) Mol Cell 51: 360
- PubMed: 23932717 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2013.07.011
- Primary Citation Related Structures:
4BWK, 4BWP, 4BWX - PubMed Abstract:
The PAN2-PAN3 deadenylase complex functions in general and miRNA-mediated mRNA degradation and is specifically recruited to miRNA targets by GW182/TNRC6 proteins. We describe the PAN3 adaptor protein crystal structure that, unexpectedly, forms intertwined and asymmetric homodimers. Dimerization is mediated by a coiled coil that links an N-terminal pseudokinase to a C-terminal knob domain. The PAN3 pseudokinase binds ATP, and this function is required for mRNA degradation in vivo. We further identified conserved surfaces required for mRNA degradation, including the binding surface for the PAN2 deadenylase on the knob domain. The most remarkable structural feature is the presence of a tryptophan-binding pocket at the dimer interface, which mediates binding to TNRC6C in human cells. Together, our data reveal the structural basis for the interaction of PAN3 with PAN2 and the recruitment of the PAN2-PAN3 complex to miRNA targets by TNRC6 proteins.
- Department of Biochemistry, Max Planck Institute for Developmental Biology, Spemannstrasse 35, 72076 Tübingen, Germany.
Organizational Affiliation: