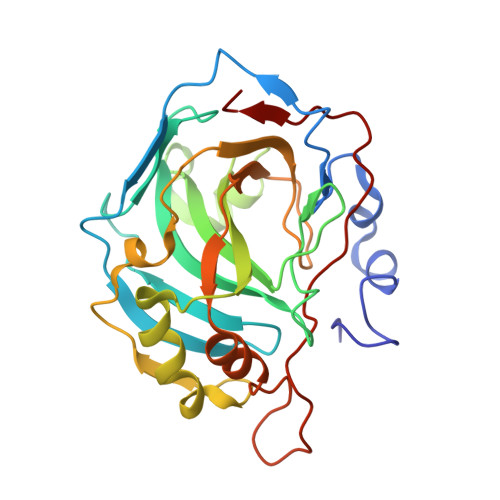
Sulfocoumarins (1,2-Benzoxathiine-2,2-Dioxides): A Class of Potent and Isoform-Selective Inhibitors of Tumor-Associated Carbonic Anhydrases.
Tars, K., Vullo, D., Kazaks, A., Leitans, J., Lends, A., Grandane, A., Zalubovskis, R., Scozzafava, A., Supuran, C.T.(2013) J Med Chem 56: 293
- PubMed: 23241068 Search on PubMed
- DOI: https://doi.org/10.1021/jm301625s
- Primary Citation Related Structures:
4BCW - PubMed Abstract:
Coumarins were recently shown to constitute a novel class of mechanism-based carbonic anhydrase (CA, EC 4.2.1.1) inhibitors. We demonstrate that sulfocoumarins (1,2-benzoxathiine 2,2-dioxides) possess a similar mechanism of action, acting as effective CA inhibitors. The sulfocoumarins were hydrolyzed by the esterase CA activity to 2-hydroxyphenyl-vinylsulfonic acids, which thereafter bind to the enzyme in a region rarely occupied by other classes of inhibitors. The X-ray structure of one of these compounds in adduct with a modified CA II enzyme possessing two amino acid residues from the CA IX active site, allowed us to decipher the inhibition mechanism. The sulfonic acid was observed anchored to the zinc-coordinated water molecule, making favorable interactions with Thr200 and Pro201. Some other sulfocoumarins incorporating substituted-1,2,3-triazole moieties were prepared by using click chemistry and showed low nanomolar inhibitory action against the tumor-associated isoforms CA IX and XII, being less effective against the cytosolic CA I and II.
- Biomedical Research and Study Center, Ratsupites 1, LV 1067 Riga, Latvia.
Organizational Affiliation: