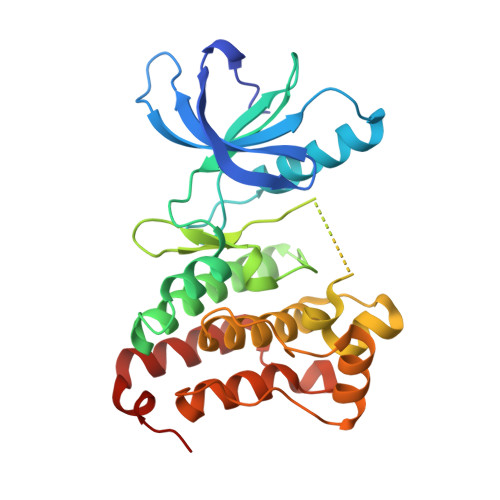
The Design, Synthesis, and Biological Evaluation of Potent Receptor Tyrosine Kinase Inhibitors.
Kim, M.H., Tsuhako, A.L., Co, E.W., Aftab, D.T., Bentzien, F., Chen, J., Cheng, W., Engst, S., Goon, L., Klein, R.R., Le, D.T., Mac, M., Parks, J.J., Qian, F., Rodriquez, M., Stout, T.J., Till, J.H., Won, K.A., Wu, X., Michael Yakes, F., Yu, P., Zhang, W., Zhao, Y., Lamb, P., Nuss, J.M., Xu, W.(2012) Bioorg Med Chem Lett 22: 4979
- PubMed: 22765894 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2012.06.029
- Primary Citation Related Structures:
4AW5 - PubMed Abstract:
Variously substituted indolin-2-ones were synthesized and evaluated for activity against KDR, Flt-1, FGFR-1 and PDGFR. Extension at the 5-position of the oxindole ring with ethyl piperidine (compound 7i) proved to be the most beneficial for attaining both biochemical and cellular potencies. Further optimization of 7i to balance biochemical and cellular potencies with favorable ADME/ PK properties led to the identification of 8h, a compound with a clean CYP profile, acceptable pharmacokinetic and toxicity profiles, and robust efficacy in multiple xenograft tumor models.
- Exelixis, 210 E. Grand Avenue, South San Francisco, CA 94080, USA.
Organizational Affiliation: