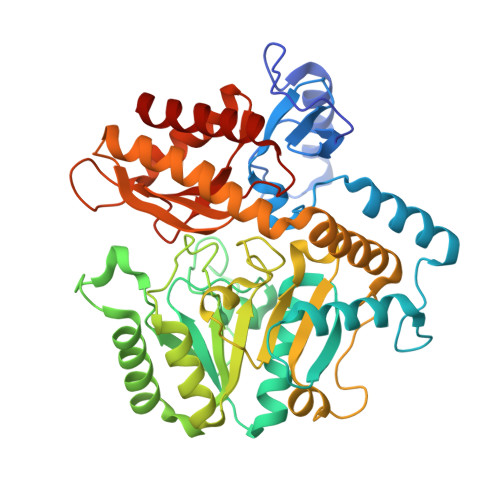
Structures of a Gamma-Aminobutyrate (Gaba) Transaminase from the S-Triazine-Degrading Organism Arthrobacter Aurescens Tc1 in Complex with Plp and with its External Aldimine Plp- Gaba Adduct.
Bruce, H., Nguyen Tuan, A., Mangas Sanchez, J., Leese, C., Hopwood, J., Hyde, R., Hart, S., Turkenburg, J.P., Grogan, G.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 1175
- PubMed: 23027742 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112030023
- Primary Citation Related Structures:
4ATP, 4ATQ - PubMed Abstract:
Two complex structures of the γ-aminobutyrate (GABA) transaminase A1R958 from Arthrobacter aurescens TC1 are presented. The first, determined to a resolution of 2.80 Å, features the internal aldimine formed by reaction between the ℇ-amino group of Lys295 and the cofactor pyridoxal phosphate (PLP); the second, determined to a resolution of 2.75 Å, features the external aldimine adduct formed between PLP and GABA in the first half-reaction. This is the first structure of a microbial GABA transaminase in complex with its natural external aldimine and reveals the molecular determinants of GABA binding in this enzyme.
- York Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5DD, England.
Organizational Affiliation: