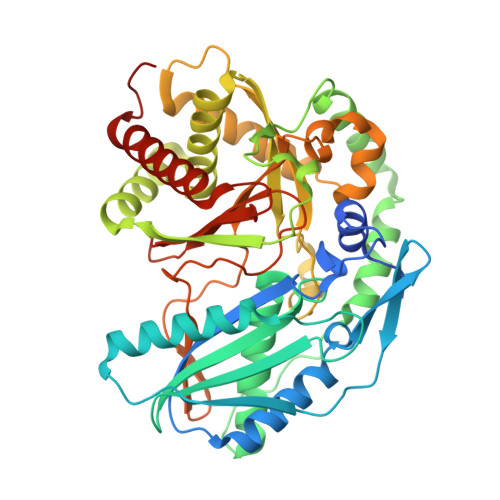
Crystal structure of SgcC5 protein from Streptomyces globisporus
Michalska, K., Bigelow, L., Jedrzejczak, R., Babnigg, G., Lohman, J., Ma, M., Rudolf, J., Chang, C.-Y., Shen, B., Joachimiak, A., Midwest Center for Structural Genomics (MCSG), Enzyme Discovery for Natural Product Biosynthesis (NatPro)To be published.