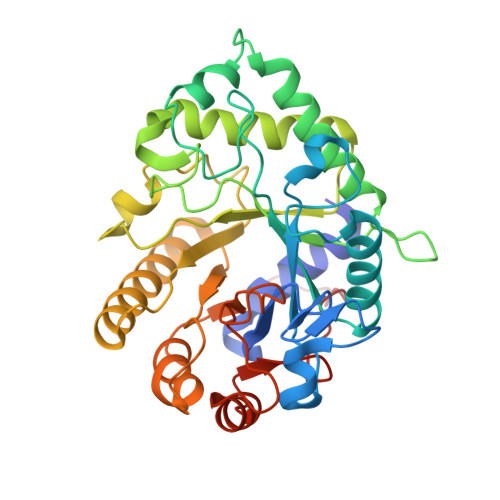
Galactomannan Catabolism Conferred by a Polysaccharide Utilization Locus of Bacteroides ovatus: ENZYME SYNERGY AND CRYSTAL STRUCTURE OF A beta-MANNANASE.
Bagenholm, V., Reddy, S.K., Bouraoui, H., Morrill, J., Kulcinskaja, E., Bahr, C.M., Aurelius, O., Rogers, T., Xiao, Y., Logan, D.T., Martens, E.C., Koropatkin, N.M., Stalbrand, H.(2017) J Biological Chem 292: 229-243
- PubMed: 27872187
- DOI: https://doi.org/10.1074/jbc.M116.746438
- Primary Citation Related Structures:
4ZXO - PubMed Abstract:
A recently identified polysaccharide utilization locus (PUL) from Bacteroides ovatus ATCC 8483 is transcriptionally up-regulated during growth on galacto- and glucomannans. It encodes two glycoside hydrolase family 26 (GH26) β-mannanases, BoMan26A and BoMan26B, and a GH36 α-galactosidase, BoGal36A. The PUL also includes two glycan-binding proteins, confirmed by β-mannan affinity electrophoresis. When this PUL was deleted, B. ovatus was no longer able to grow on locust bean galactomannan. BoMan26A primarily formed mannobiose from mannan polysaccharides. BoMan26B had higher activity on galactomannan with a high degree of galactosyl substitution and was shown to be endo-acting generating a more diverse mixture of oligosaccharides, including mannobiose. Of the two β-mannanases, only BoMan26B hydrolyzed galactoglucomannan. A crystal structure of BoMan26A revealed a similar structure to the exo-mannobiohydrolase CjMan26C from Cellvibrio japonicus, with a conserved glycone region (-1 and -2 subsites), including a conserved loop closing the active site beyond subsite -2. Analysis of cellular location by immunolabeling and fluorescence microscopy suggests that BoMan26B is surface-exposed and associated with the outer membrane, although BoMan26A and BoGal36A are likely periplasmic. In light of the cellular location and the biochemical properties of the two characterized β-mannanases, we propose a scheme of sequential action by the glycoside hydrolases encoded by the β-mannan PUL and involved in the β-mannan utilization pathway in B. ovatus. The outer membrane-associated BoMan26B initially acts on the polysaccharide galactomannan, producing comparably large oligosaccharide fragments. Galactomanno-oligosaccharides are further processed in the periplasm, degalactosylated by BoGal36A, and subsequently hydrolyzed into mainly mannobiose by the β-mannanase BoMan26A.
- From the Department of Biochemistry and Structural Biology, Lund University P. O. Box 124, S-221 00 Lund, Sweden and.
Organizational Affiliation: