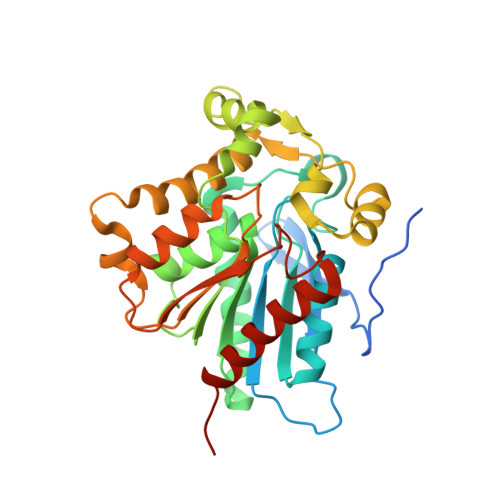
Crystal structure of the Saccharomyces cerevisiae monoglyceride lipase Yju3p.
Aschauer, P., Rengachari, S., Lichtenegger, J., Schittmayer, M., Das, K.M., Mayer, N., Breinbauer, R., Birner-Gruenberger, R., Gruber, C.C., Zimmermann, R., Gruber, K., Oberer, M.(2016) Biochim Biophys Acta 1861: 462-470
- PubMed: 26869448 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbalip.2016.02.005
- Primary Citation Related Structures:
4ZWN, 4ZXF - PubMed Abstract:
Monoglyceride lipases (MGLs) are a group of α/β-hydrolases that catalyze the hydrolysis of monoglycerides (MGs) into free fatty acids and glycerol. This reaction serves different physiological functions, namely in the last step of phospholipid and triglyceride degradation, in mammalian endocannabinoid and arachidonic acid metabolism, and in detoxification processes in microbes. Previous crystal structures of MGLs from humans and bacteria revealed conformational plasticity in the cap region of this protein and gave insight into substrate binding. In this study, we present the structure of a MGL from Saccharomyces cerevisiae called Yju3p in its free form and in complex with a covalently bound substrate analog mimicking the tetrahedral intermediate of MG hydrolysis. These structures reveal a high conservation of the overall shape of the MGL cap region and also provide evidence for conformational changes in the cap of Yju3p. The complex structure reveals that, despite the high structural similarity, Yju3p seems to have an additional opening to the substrate binding pocket at a different position compared to human and bacterial MGL. Substrate specificities towards MGs with saturated and unsaturated alkyl chains of different lengths were tested and revealed highest activity towards MG containing a C18:1 fatty acid.
- Institute of Molecular Biosciences, University of Graz, Humboldtstraße 50/3, 8010 Graz, Austria.
Organizational Affiliation: