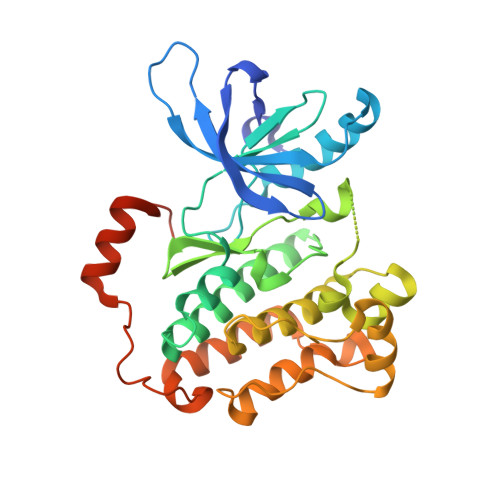
Ibrutinib Selectively and Irreversibly Targets EGFR-mutant non-Small Cell Lung Cancer Cells
Wu, H., Wang, A., Zhang, W., Wang, B., Wang, B., Yan, X.E., Chen, C., Hu, C., Ye, Z., Zhao, Z., Wang, L., Li, X., Yu, K., Liu, J., Wu, J., Wang, J., Wang, C., Weisberg, E.L., Liu, J., Gray, N.S., Chen, L., Yun, C.H., Liu, Q.To be published.