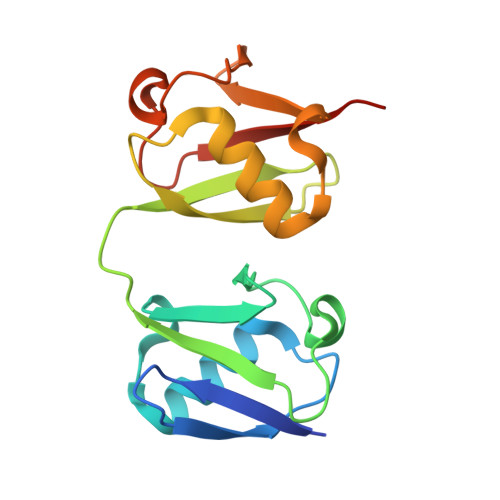
New conformations of linear polyubiquitin chains from crystallographic and solution-scattering studies expand the conformational space of polyubiquitin.
Thach, T.T., Shin, D., Han, S., Lee, S.(2016) Acta Crystallogr D Struct Biol 72: 524-535
- PubMed: 27050132 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798316001510
- Primary Citation Related Structures:
4ZQS - PubMed Abstract:
The conformational flexibility of linkage-specific polyubiquitin chains enables ubiquitylated proteins and their receptors to be involved in a variety of cellular processes. Linear or Met1-linked polyubiquitin chains, associated with nondegradational cellular signalling pathways, have been known to adopt multiple conformations from compact to extended conformations. However, the extent of such conformational flexibility remains open. Here, the crystal structure of linear Ub2 was determined in a more compact conformation than that of the previously known structure (PDB entry 3axc). The two structures differ significantly from each other, as shown by an r.m.s.d. between C(α) atoms of 3.1 Å. The compactness of the linear Ub2 structure in comparison with PDB entry 3axc is supported by smaller values of the radius of gyration (Rg; 18 versus 18.9 Å) and the maximum interatomic distance (Dmax; 55.5 versus 57.8 Å). Extra intramolecular hydrogen bonds formed among polar residues between the distal and proximal ubiquitin moieties seem to contribute to stabilization of the compact conformation of linear Ub2. An ensemble of three semi-extended and extended conformations of linear Ub2 was also observed by small-angle X-ray scattering (SAXS) analysis in solution. In addition, the conformational heterogeneity in linear polyubiquitin chains is clearly manifested by SAXS analyses of linear Ub3 and Ub4: at least three distinct solution conformations are observed in each chain, with the linear Ub3 conformations being compact. The results expand the extent of conformational space of linear polyubiquitin chains and suggest that changes in the conformational ensemble may be pivotal in mediating multiple signalling pathways.
- Department of Biological Sciences, Sungkyunkwan University, 2066 Seobu-ro, Suwon, Gyeonggi 16419, Republic of Korea.
Organizational Affiliation: